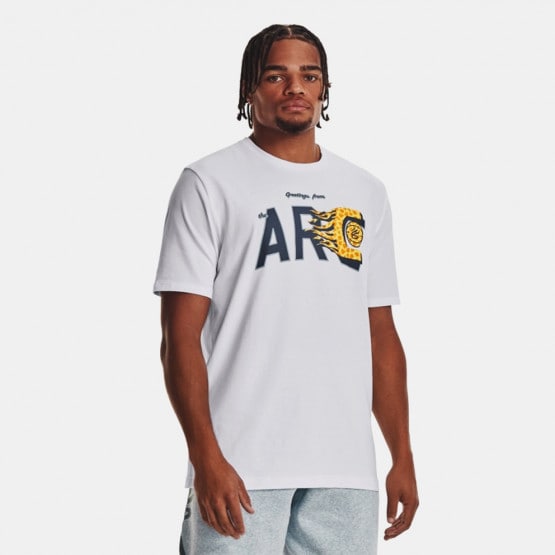
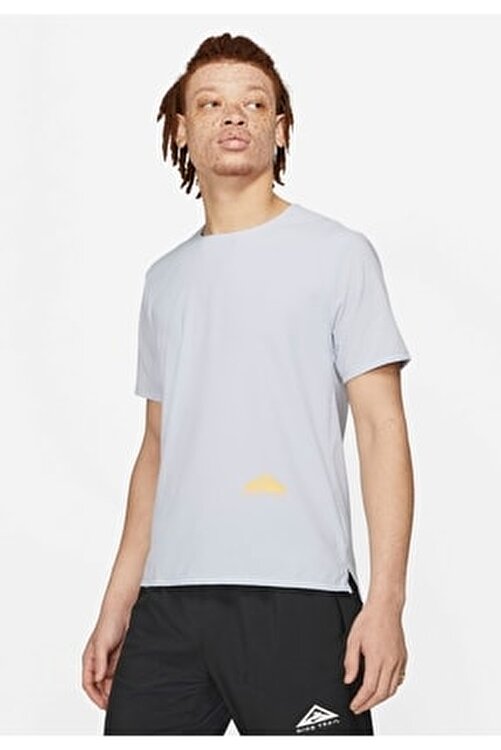
Arvind Sport | Training & CrossFit Clothes. Discover the Men's & Women's & Kids' Collection | Offers | Stock (8), Autry round neck short-sleeved T- shirt Blu

楽天市場】【公式】アディダス adidas 返品可 Designed for Gameday アンセムジャケット スポーツウェア メンズ ウェア・服 トップス ジャージ 白 ホワイト HT3402 : adidas Online Shop 楽天市場店

Gottliebpaludan Sport, Προσφορές, Στοκ, Βρες adidas Γυναικεία Παπούτσια, Ρούχα & Αξεσουάρ. adidas Performance | adidas Originals | για προπόνηση και για τρέξιμο και για καθημερινό, yeezy sneakers waverunners shoes clearance list