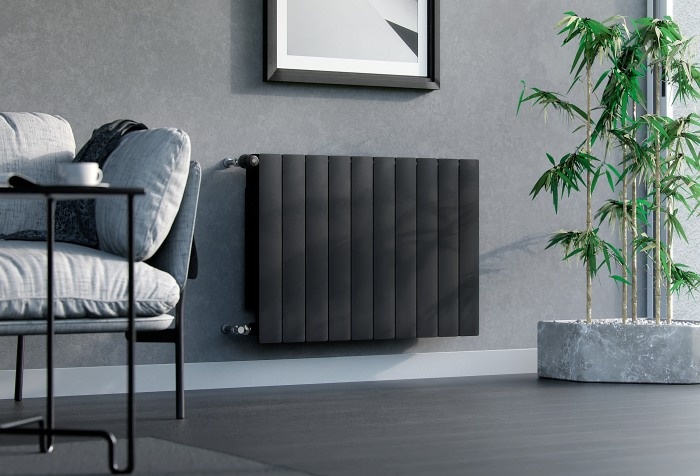
Klasyka sprawdzona przez miliony – dlaczego grzejnik aluminiowy to faktycznie dobry wybór? | RynekInstalacyjny.pl
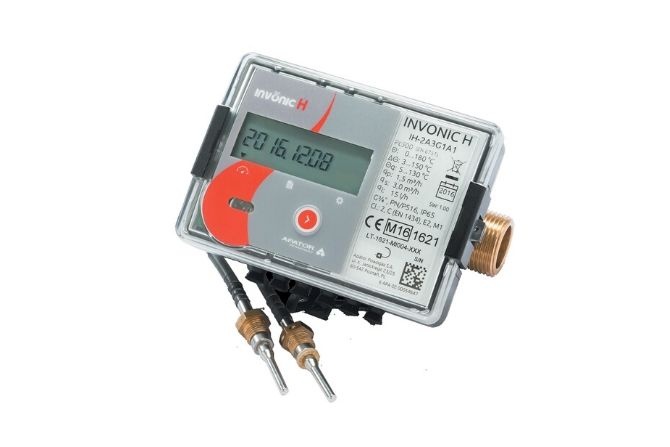

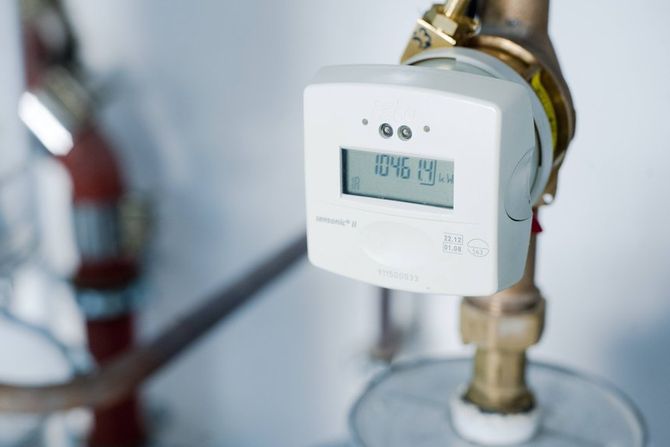
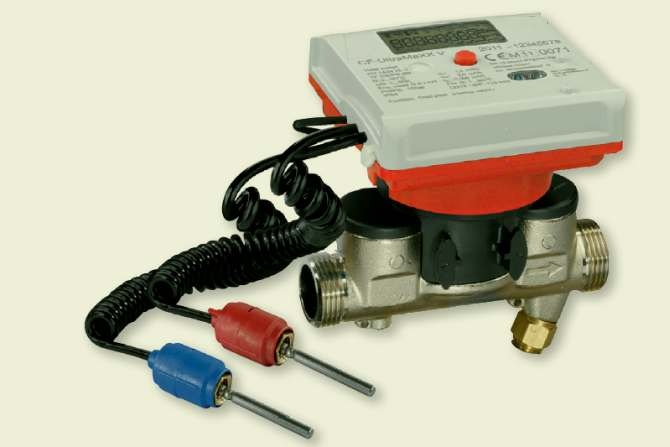
Ciepłomierze indywidualne – wybrane prawne i ekonomiczne aspekty doboru, montażu i eksploatacji | RynekInstalacyjny.pl

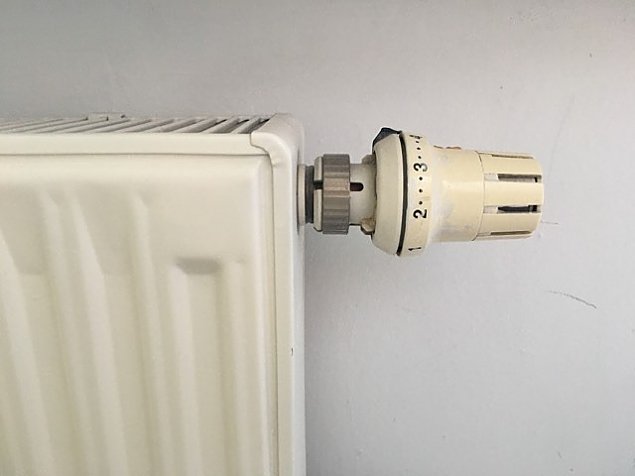
WHY IS THE HEATER COLD AT THE BOTTOM? The radiator is only hot at the top! Heating tutorial - YouTube

WHY IS THE HEATER COLD AT THE BOTTOM? The radiator is only hot at the top! Heating tutorial - YouTube