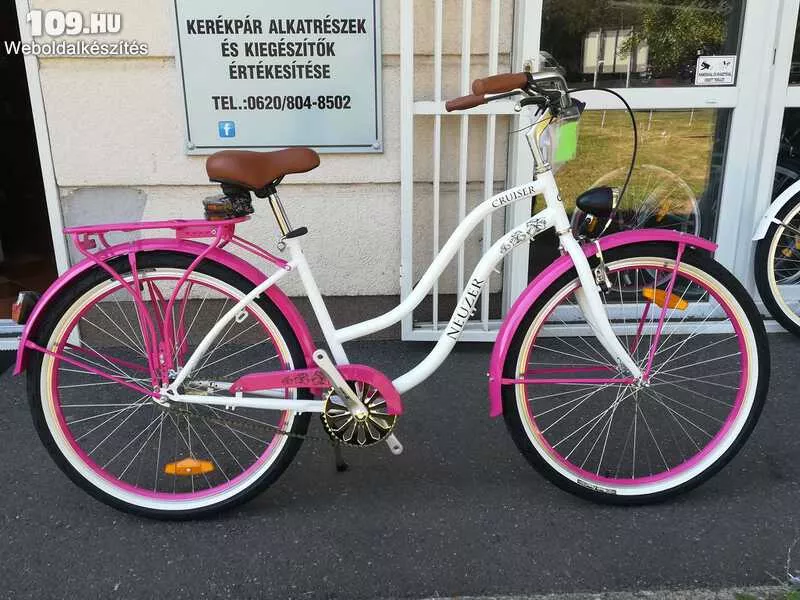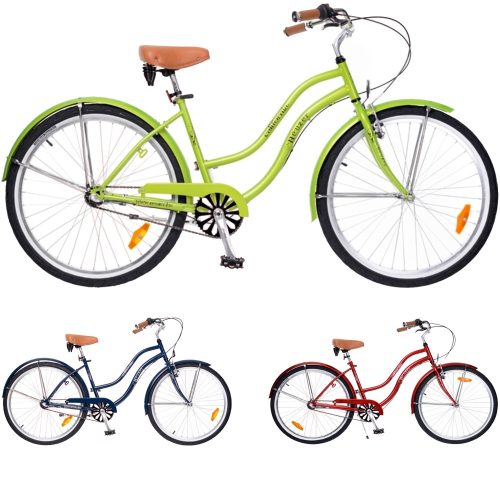
Neuzer Cruiser California N3 26” Férfi-Női kerékpár | Gondolj a nyárra! Gondolj rá milyen volt, de még inkább gondolj arra, hogy milyen lesz. Egy stílusos kerékpárral, amit nagyon jó áron még idén...

Neuzer Picnic Cruiser Női Kerékpár 26" - fehér/lila-zöld színben - Kerékpár Webshop - Webbicikli.hu - WebBicikli.hu Kerékpár Webshop
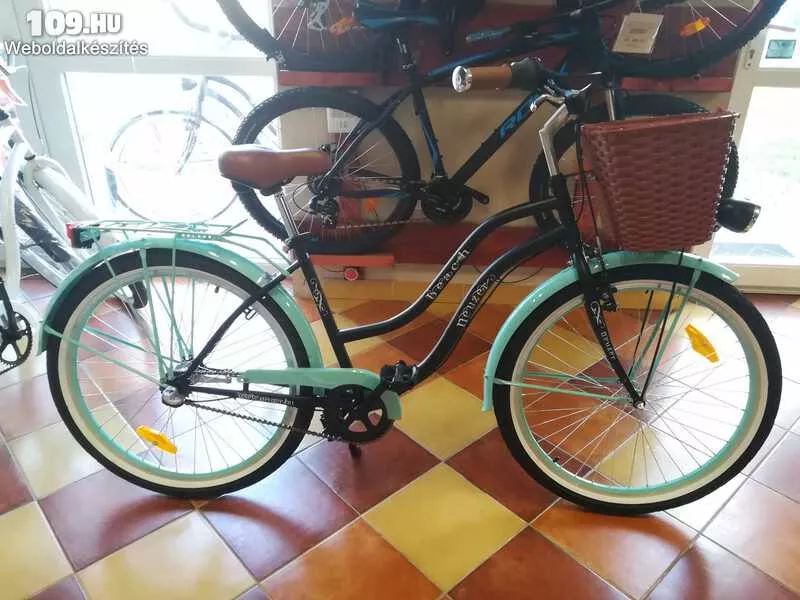

Női kerékpár Neuzer cruiser matt fekete kék egyedi agyváltós (kosár nem tartozék) - Kerékpárbolt Nyíregyháza, Nyírtura, Nyírtelek - Bringakirály Kerékpárszaküzlet és Gyorsszerviz
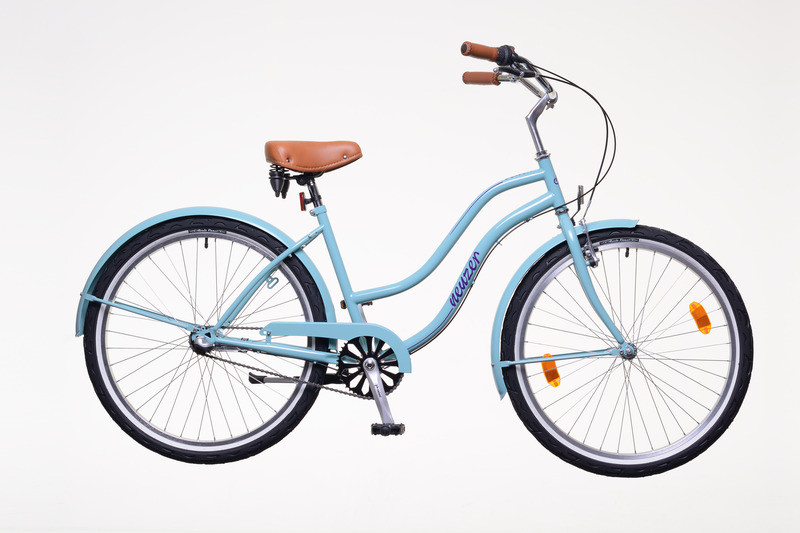

Babyblue Neuzer California női Cruiser biciklik - női kerékpár - webbicikli.hu - WebBicikli.hu Kerékpár Webshop
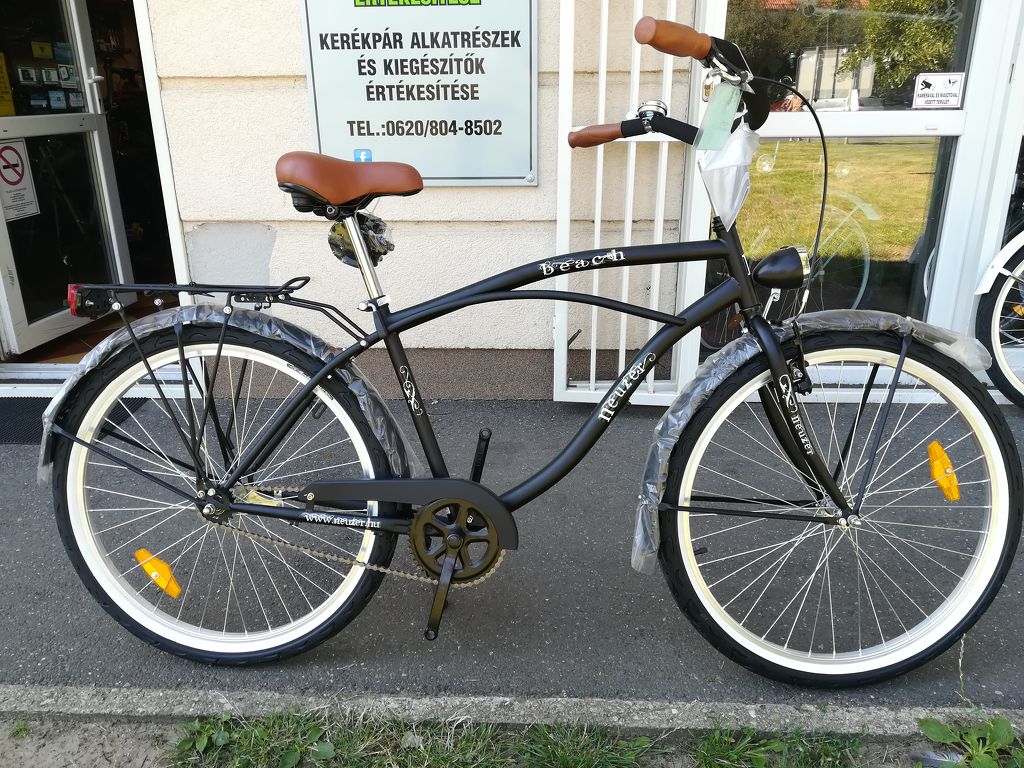

Férfi kerékpár Neuzer cruiser matt fekete felszerelt - Kerékpárbolt Nyíregyháza, Nyírtura, Nyírtelek - Bringakirály Kerékpárszaküzlet és Gyorsszerviz

Neuzer Cruiser Beach 1Seb. 26” Férfi-Női kerékpár | A nyári vízpart. Ebbena hidegben, igen, már úgy bukkan fel, mint valami vágyott meleg és kellemes emlék. Azonban lesz jövőre is nyár, hogy szép...

Őszi kerékpár Akció 8 ezer forintnyi Ajándékkal! Neuzer Cruiser Beach 1Seb. 26” Férfi-Női Női Ár: 74 900. - Akciós Ár: 74 900.- Váz... | By City Bringa | Facebook
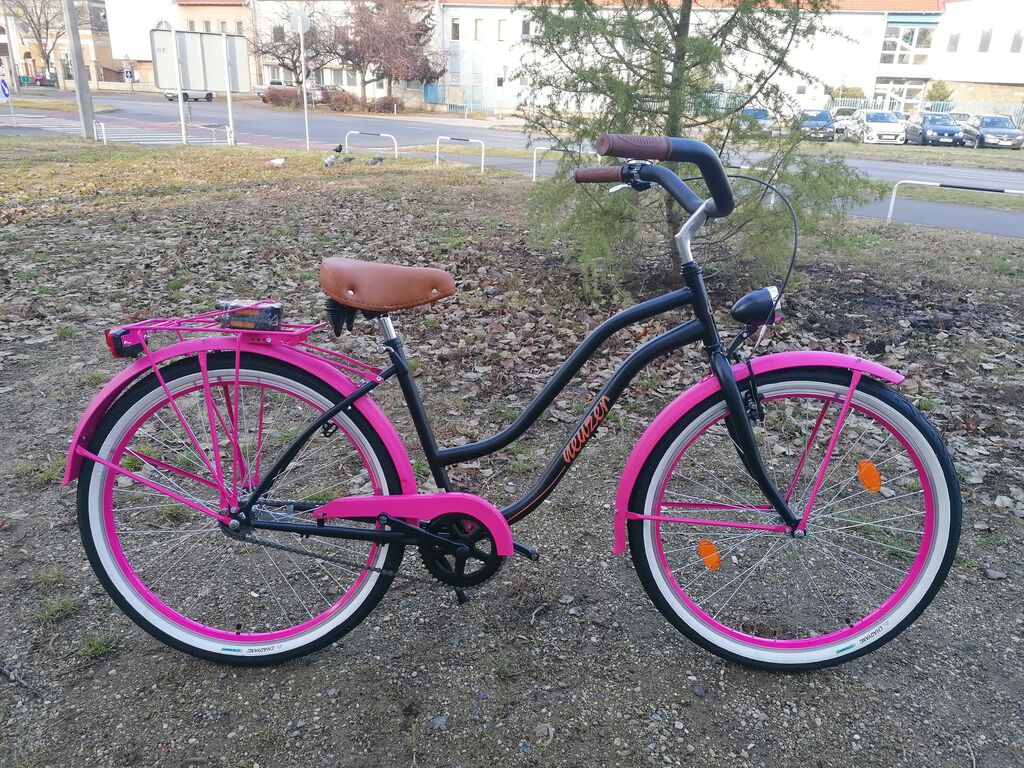

Női kerékpár Neuzer cruiser matt fekete rózsaszín felszerelt - Kerékpárbolt Nyíregyháza, Nyírtura, Nyírtelek - Bringakirály Kerékpárszaküzlet és Gyorsszerviz
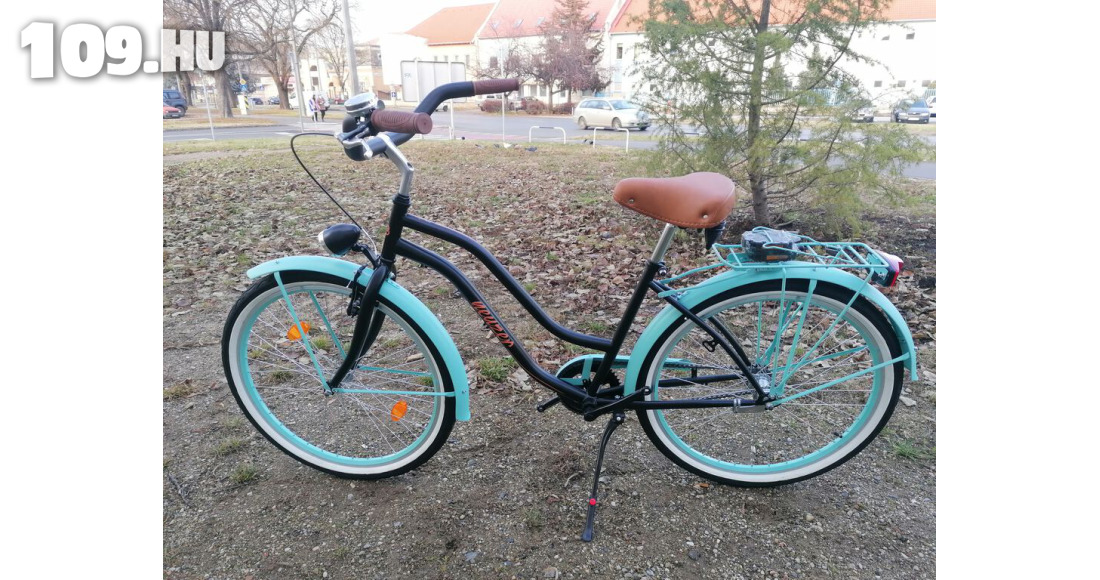

Női kerékpár Neuzer cruiser matt fekete kék felszerelt - Kerékpárbolt Nyíregyháza, Nyírtura, Nyírtelek - Bringakirály Kerékpárszaküzlet és Gyorsszerviz
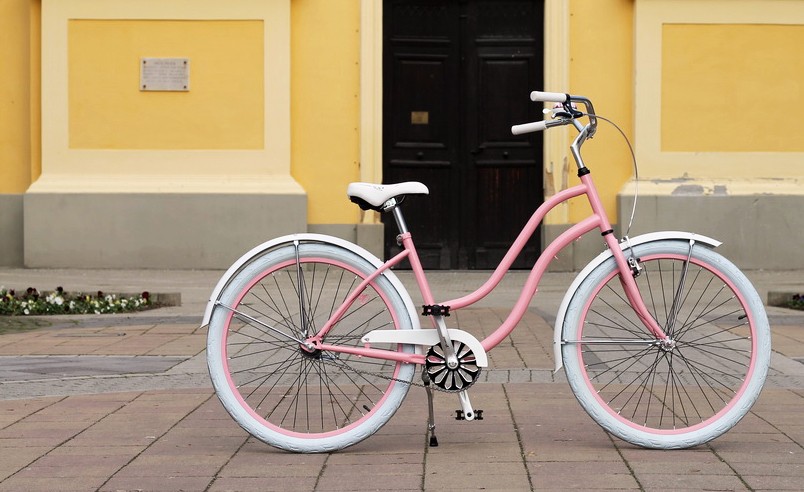
Egyedi női cruiser kerékpárok - Női Kerékpár - női kerékpárok - Cruiser kerékpár - Webbicikli.hu - WebBicikli.hu Kerékpár Webshop
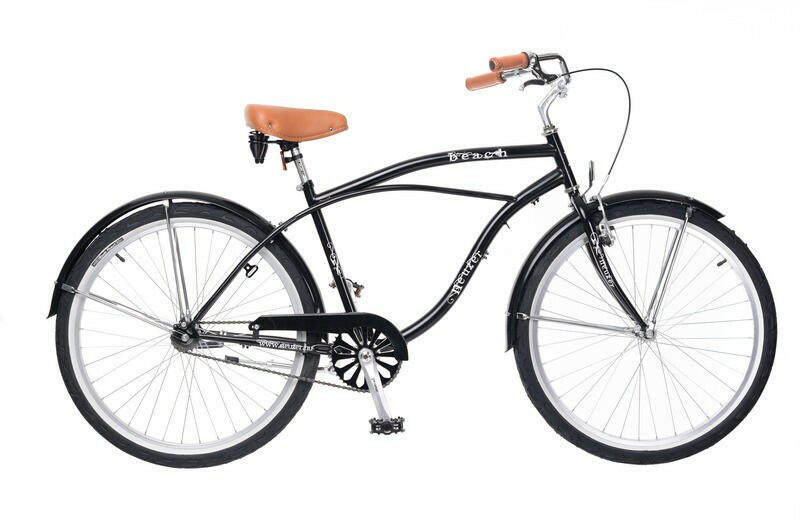

Neuzer Cruiser Beach Kerékpár árak, Kerékpár bicikli vásárlás, olcsó Kerékpárok. bringa akció, árösszehasonlító














