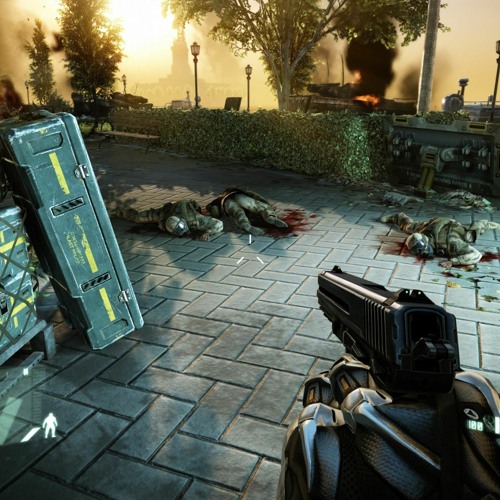
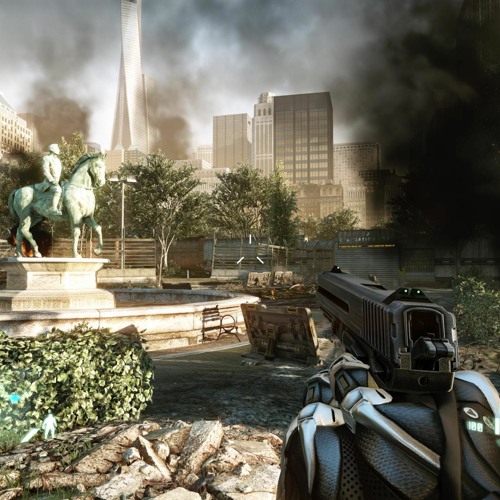
Crysis 3 Blackbox Crack Fix 2 13 2022 - Ko-fi ❤️ Where creators get support from fans through donations, memberships, shop sales and more! The original 'Buy Me a Coffee' Page.
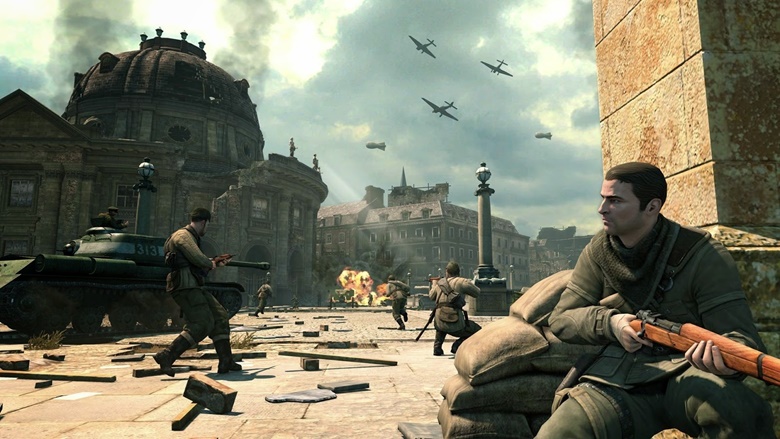

Crysis 3 Crack Fix Black Screen - Ko-fi ❤️ Where creators get support from fans through donations, memberships, shop sales and more! The original 'Buy Me a Coffee' Page.

Stream Crysis 3 Internal Reloaded Crack Fix HOT! from Simpmicressu | Listen online for free on SoundCloud