Vaistai nuo venų varikozės ir tromboflebito – Sveikas gyvenimo būdas yra pagrindinis sveikatos veiksnys

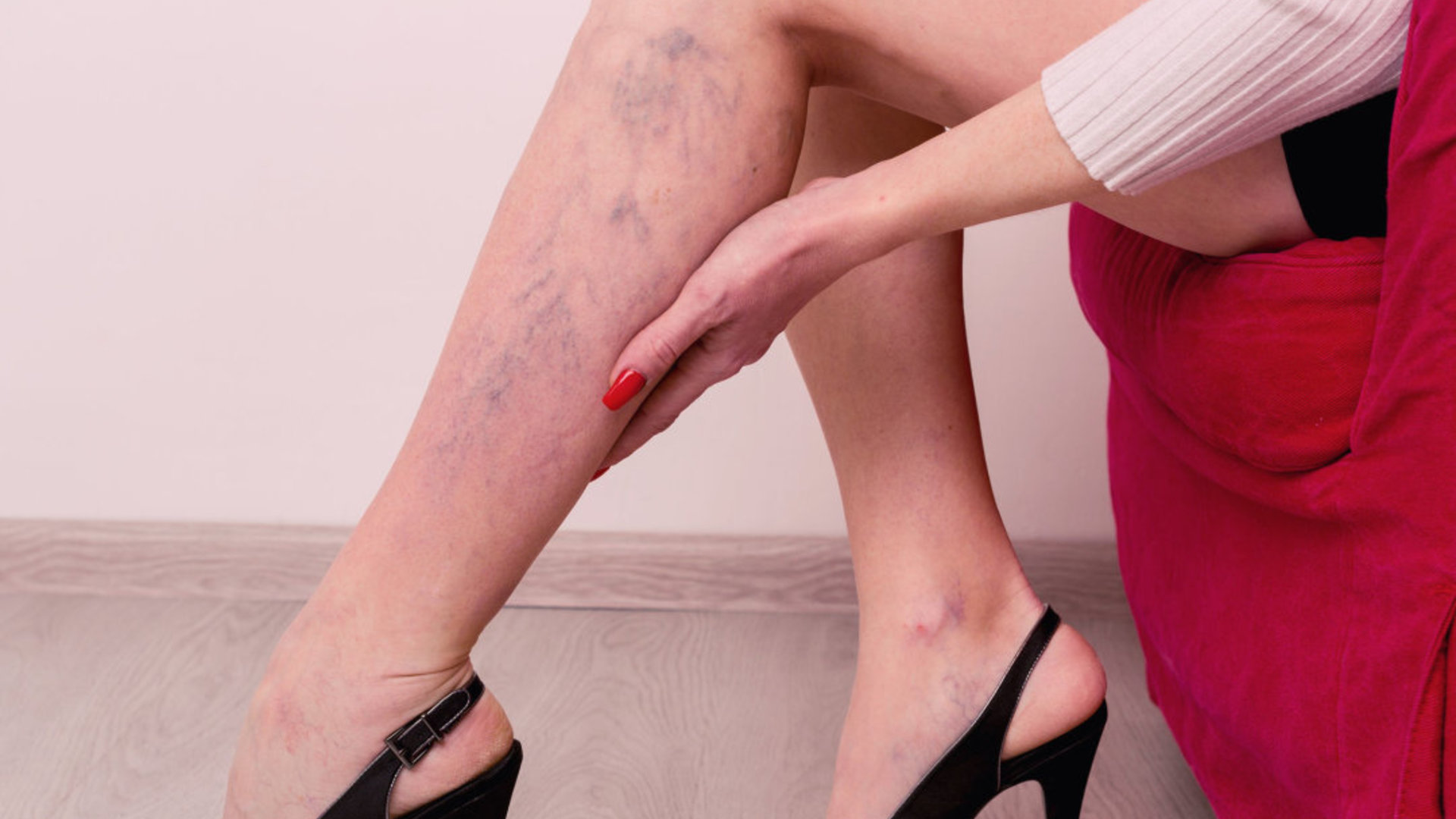
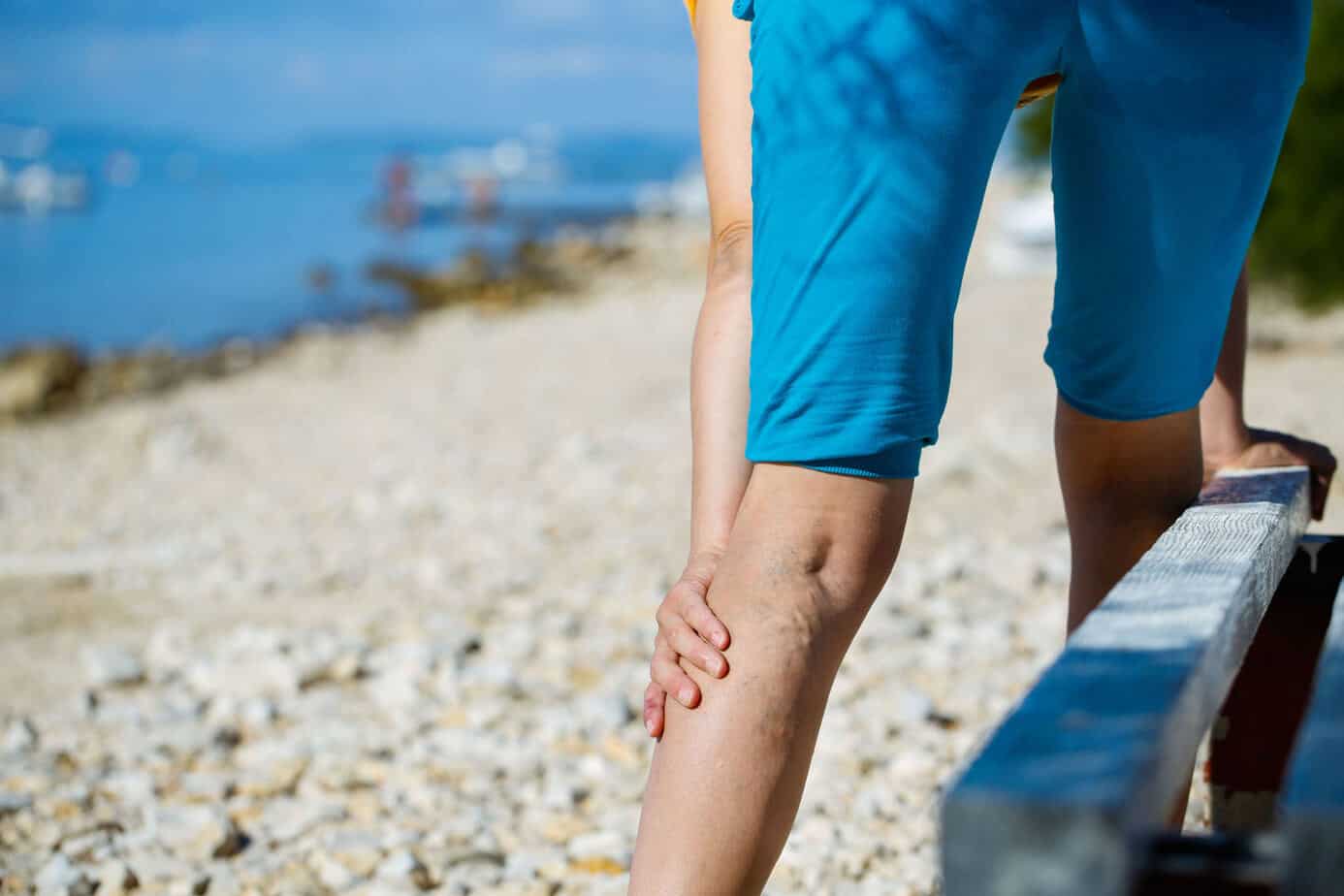
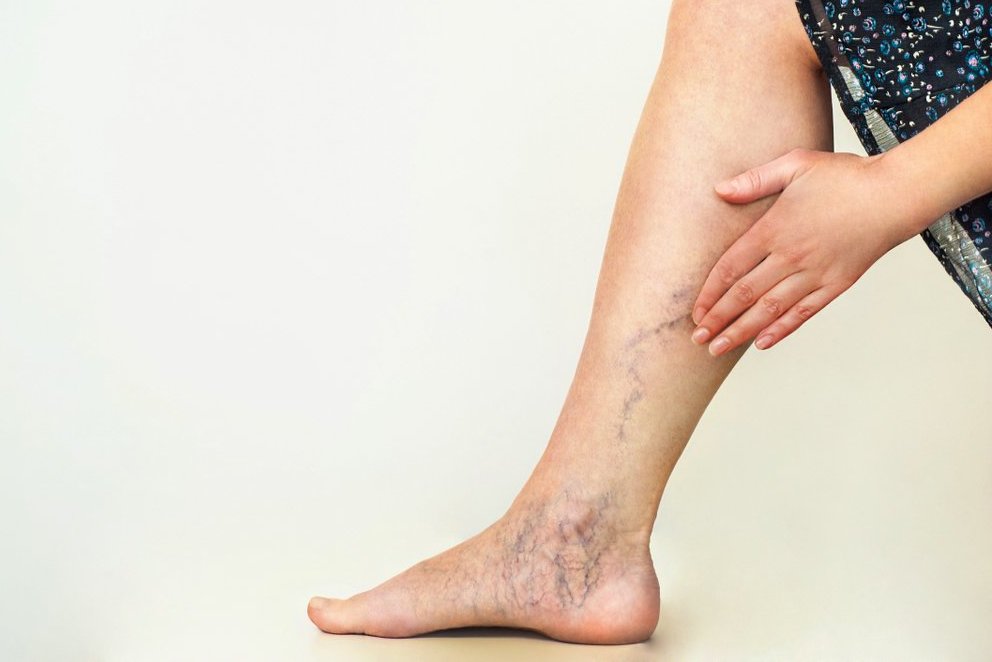
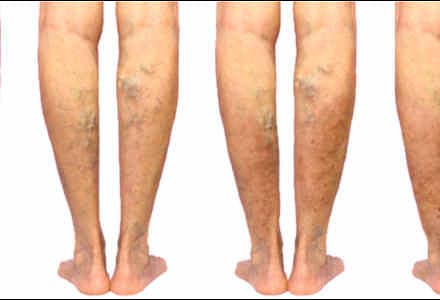
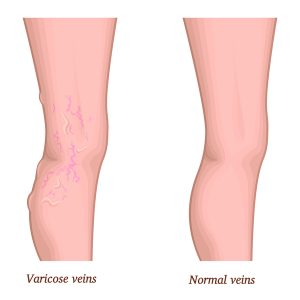
Venų varikozė ir voratinklinės venos ant kojų – Sveikas gyvenimo būdas yra pagrindinis sveikatos veiksnys
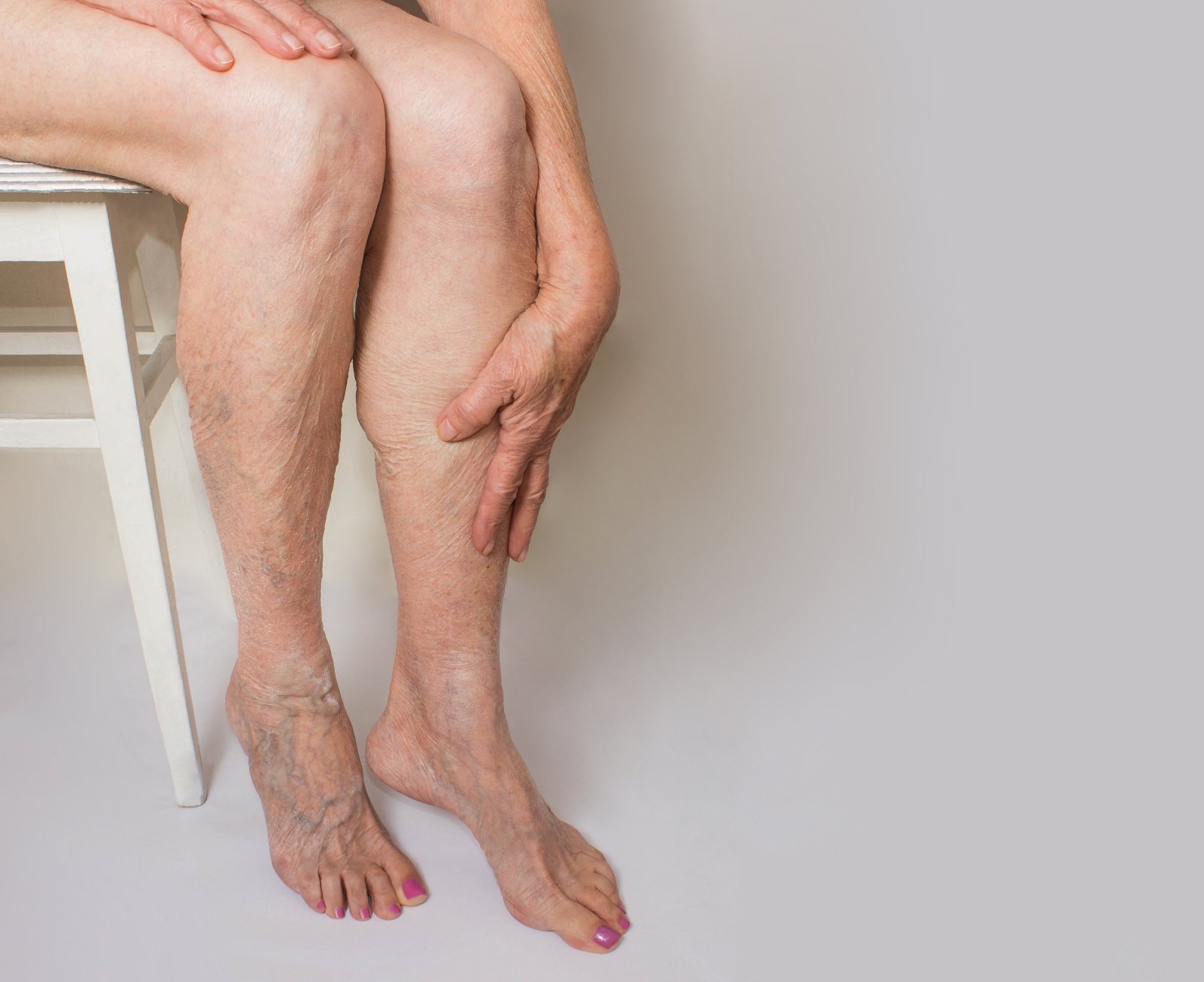
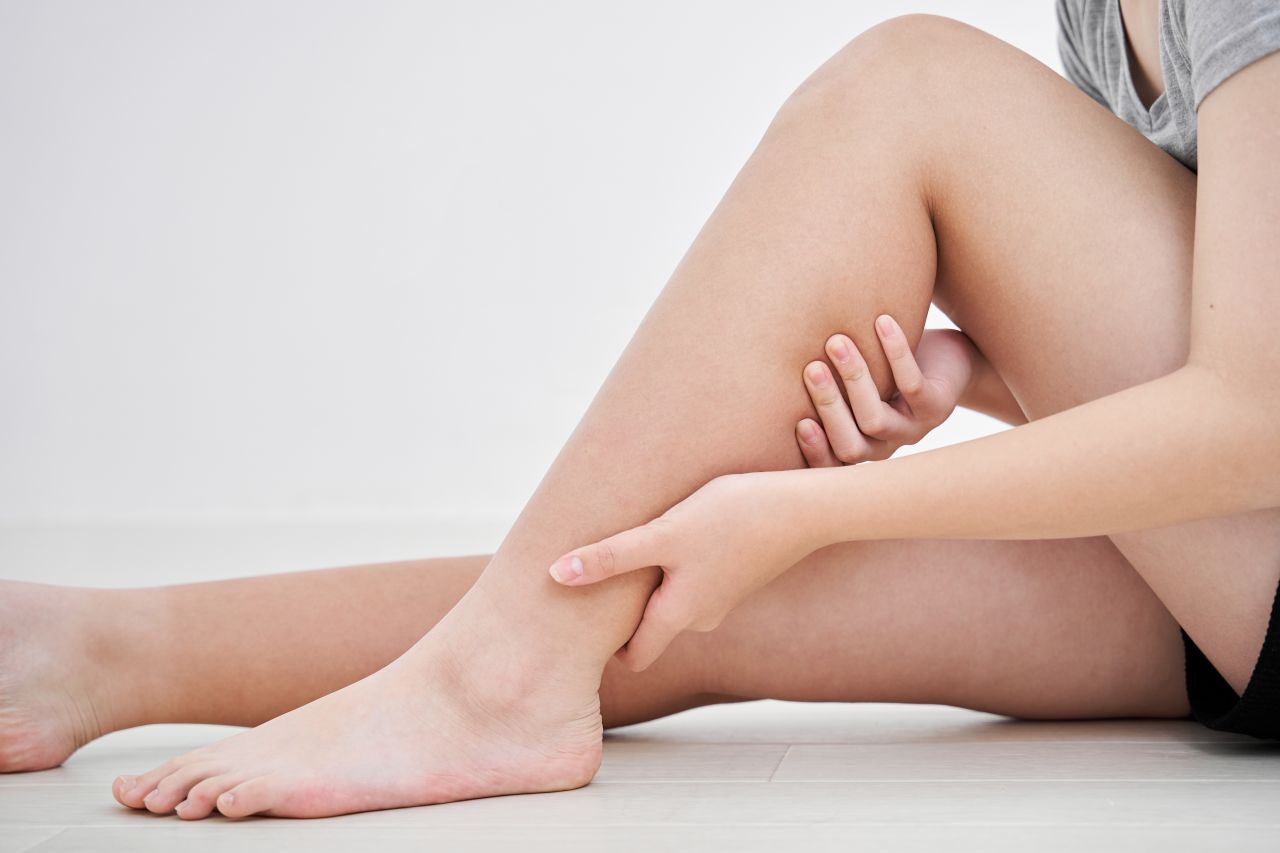
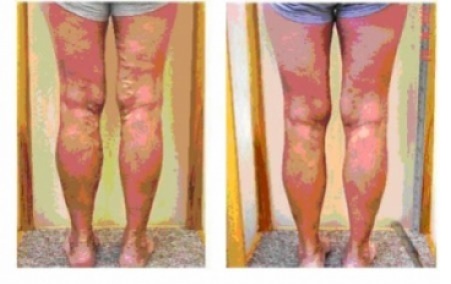
Jei laiku nesureaguosite į šiuos simptomus ant kojų, gali išsipildyti blogiausias scenarijus - DELFI