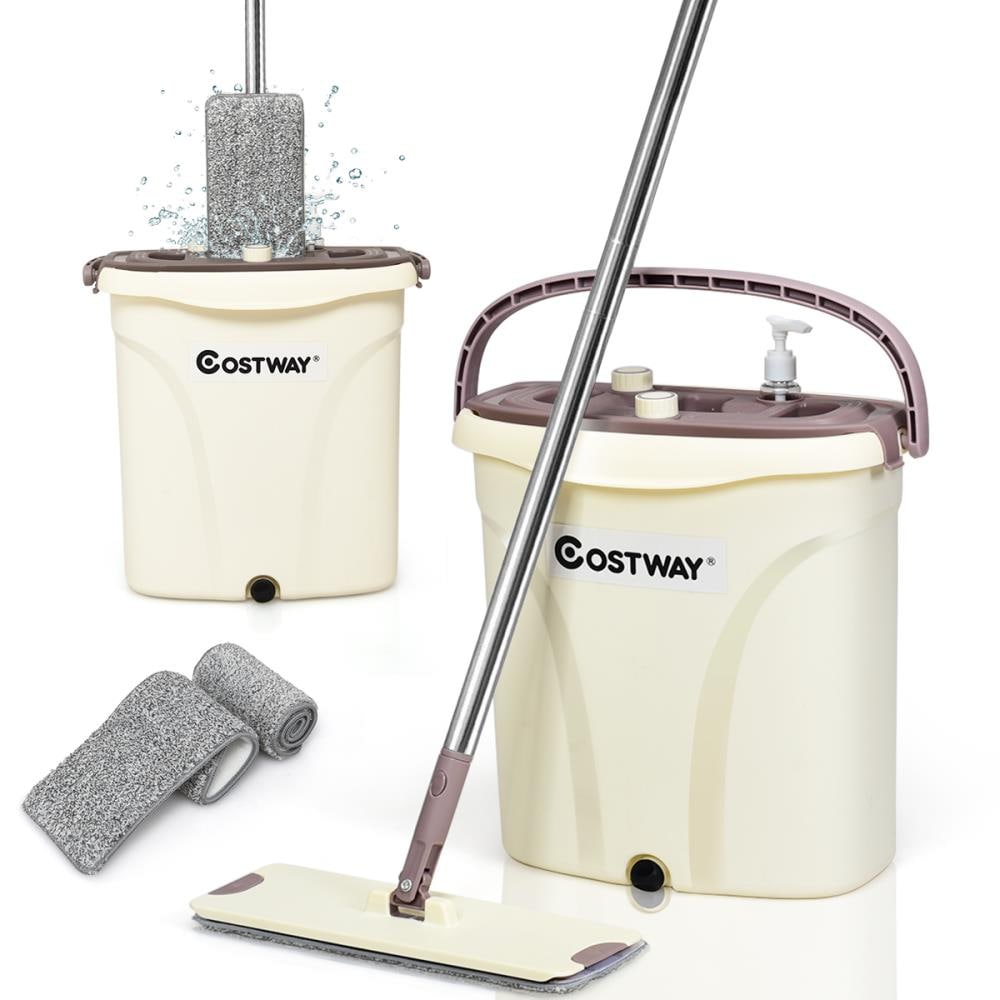
Mop Free Hand Washing New Lengthened And Thickened Lazy Wholesale Flat Mop Household Effort Saving Mop Bucket Mopping Artifact - Industrial & Scientific - Temu

360 Rotation Flat Mop Free Hand Washing Lazy Mop Floor Cleaning Microfiber Squeeze Mop Floor Clean Automatic Dehydration - Mops - AliExpress

Hand Free Flat Floor Mop And Bucket Set For Professional Home Floor Cleaning System With Washable Microfiber Pads For Hardwood - Mops - AliExpress

Goplus Flat Squeeze Mop Bucket 2 Pcs Microfiber Pad Hand-Free Wringing in the Mop Wringer Buckets department at Lowes.com
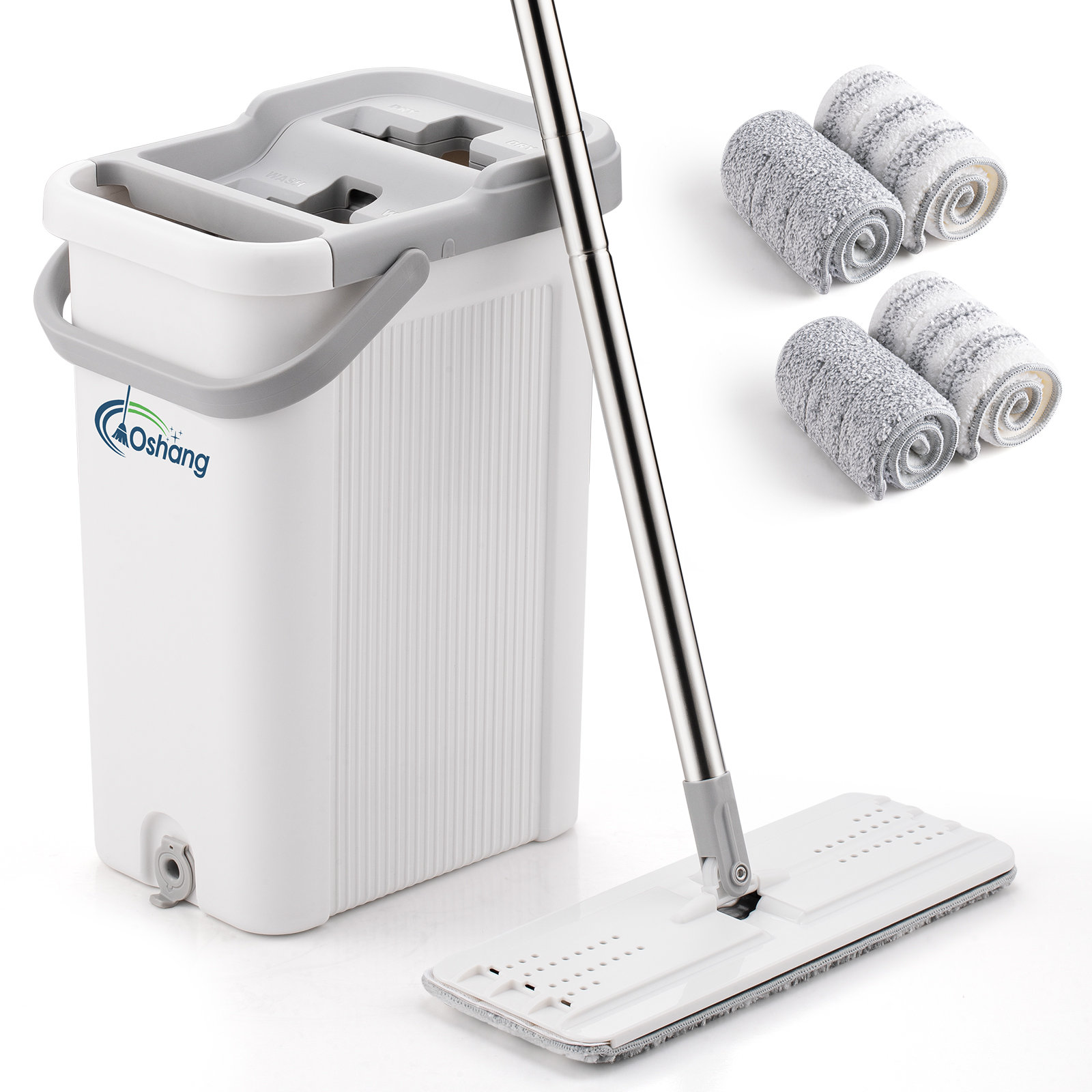

Oshang Flat Mop and Bucket OG2 - Hand-Free Floor Cleaning Mop - 4 Microfiber Mop Pads Included & Reviews | Wayfair

360 Rotation Flat Mop Free Hand Washing Lazy Mop Floor Cleaning Microfiber Squeeze Mop Floor Clean Automatic Dehydration - Mops - AliExpress

TOPMART Hands Free Flat Mop Set, Self Cleaning Wet & Dry Mop and Bucket Set, Adjustable Rectangular Mop Head & Reviews | Wayfair
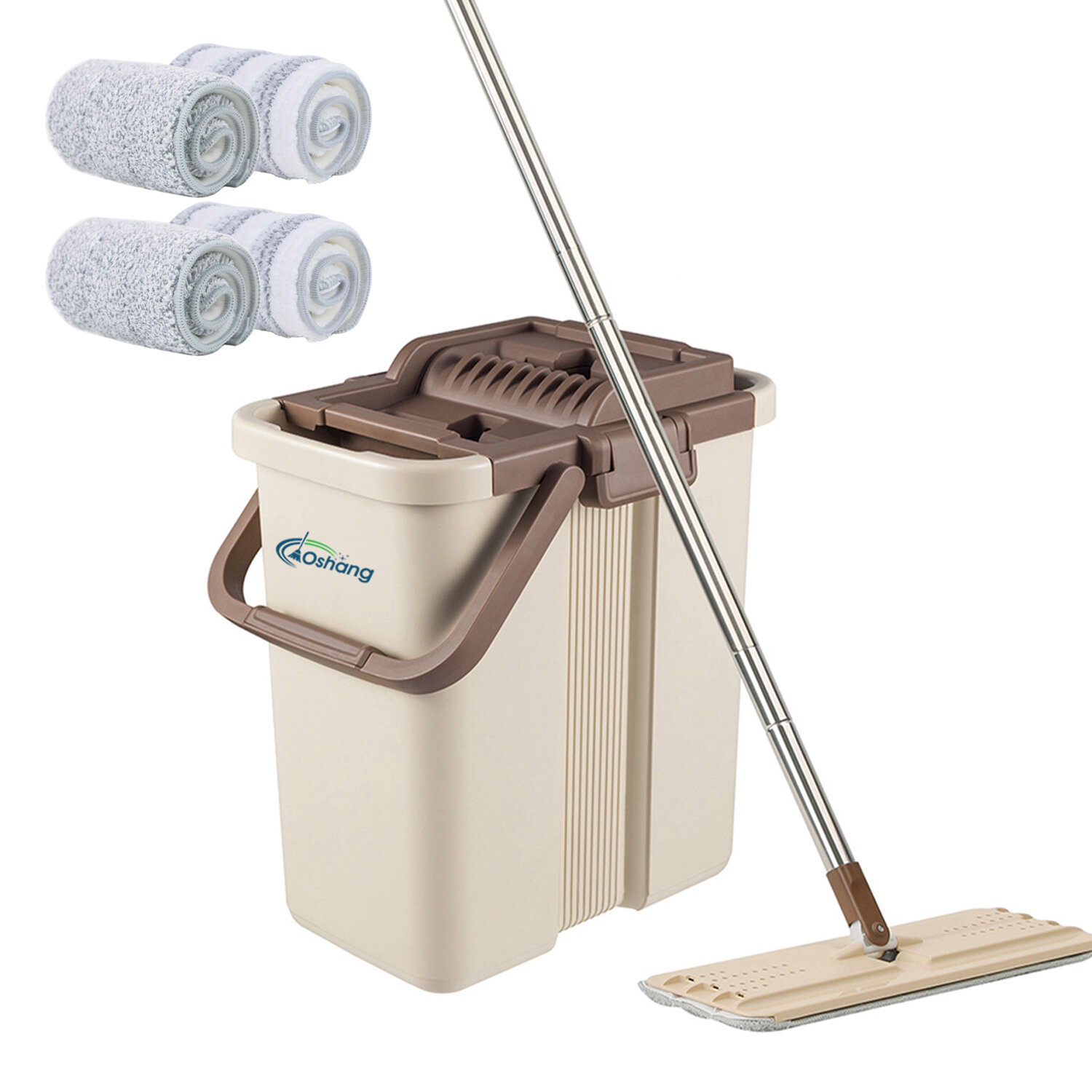

Oshang Flat Mop and Bucket OG1 - Hand-Free Floor Cleaning Mop - 4 Microfiber Mop Pads Included & Reviews | Wayfair

Flat Mop Free Hand Washing Stainless Steel Handle Spin Mop Home House Office Cleaning Tool Microfiber Pad Kitchen Floor Clean - Mops - AliExpress

Amazon.com: X3 Mop, Separates Dirty and Clean Water, 3-Chamber Design, Flat Mop and Bucket Set, Hands Free Home Floor Cleaning, 3 Reusable Microfiber Mop Pads Included : Health & Household

Amazon.com: BOSHENG Mop and Bucket with Wringer Set, Hands Free Flat Floor Mop and Bucket, 3 Washable Microfiber Pads Included, Wet and Dry Use, Home Floor Cleaning System for All Floor Types
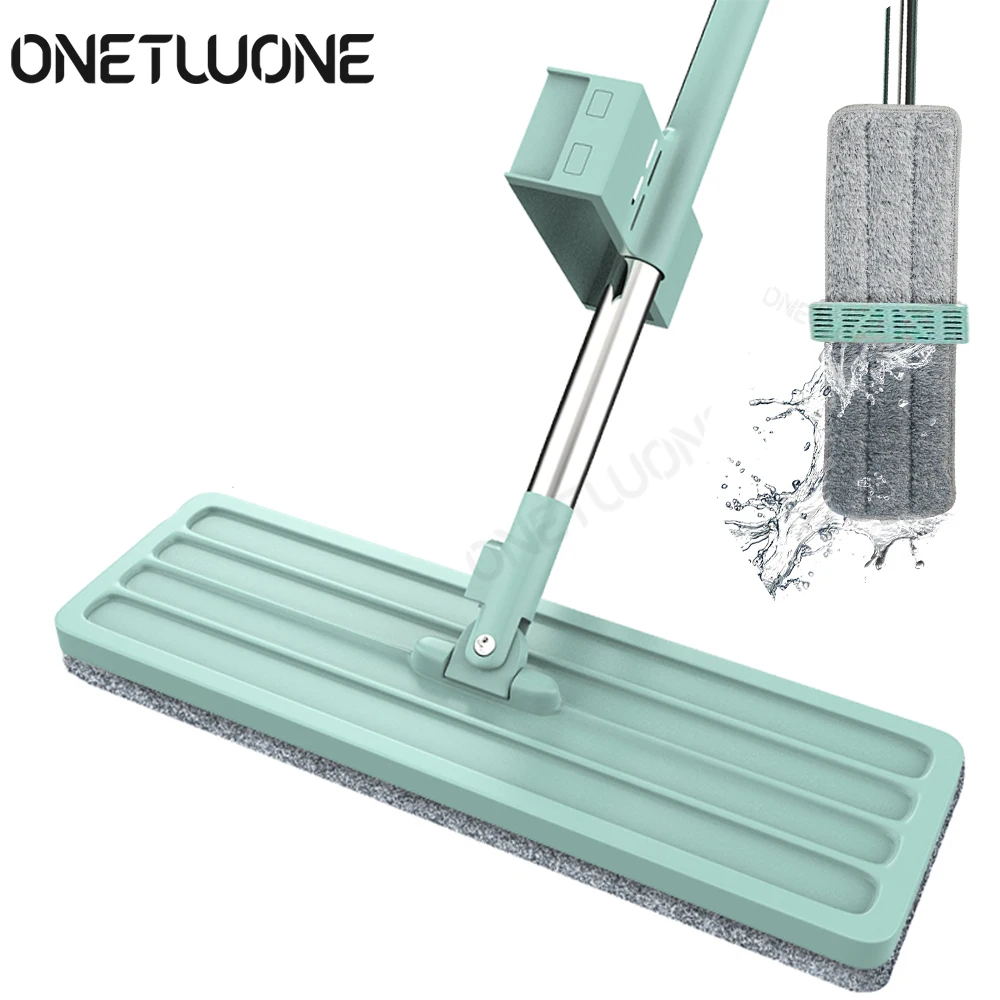

Self-wringing Magic Mop Free Hand Washing Flat Mop Automatic Spin 360 Rotating Wooden Floor Mop Cleaner Lazy Household Cleaning - Mops - AliExpress

Amazon.com: JOYMOOP Mop and Bucket with Wringer Set, Hands Free Flat Floor Mop and Bucket, with 3 Washable Microfiber Pads, Wet and Dry Use, Floor Cleaning System : Health & Household