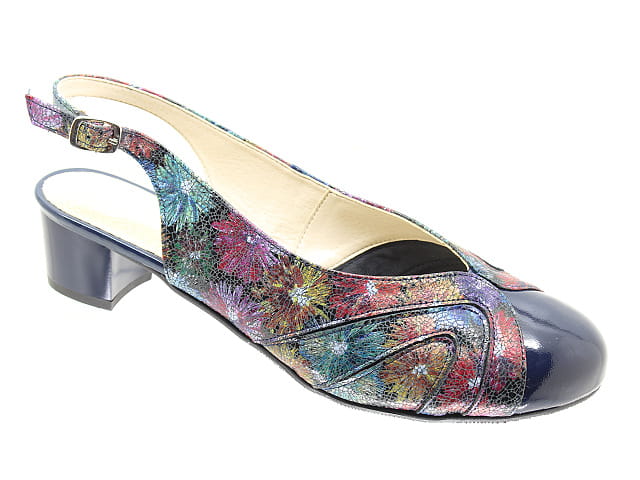
Buty xxl,Duże rozmiary butów,tęgie, szerokie, tęgość H,duże buty 41-46 ,na zamówienie, tęgie buty, Buty damskie duże rozmiary,buty na tęgą stopę,tęgość H,na tęgie stopy,na szeroką stopę,na szerokie stopy,na dużą stopę,na duże stopy,szerokie,tęgie,płaskie

Kobiety Sexy okrągłe Toe czarna wysoka podeszwa szpilki długie buty Slim Over Knee buty na cienkim obcasie duże rozmiary butów damskich|Buty za kolano| - AliExpress

DUŻE BUTY DAMSKIE VICTORIA SHOES 41 42 43 44 45 46 CZÓŁENKA BEŻOWO CZARNE . www.victoriashoes.pl na miarę i na zamówienie | victoriashoes.pl

Duże rozmiary butów damskich ,szerokie tęgie nietypowe buty XXL, na szeroką stopę,duże haluksy ,na tęgie opuchnięte chore stopy , tęgość H,K ,na zamówienie rozmiar 42,43,44,45,46, od szewca,botki,kozaki, czółenka

Buty xxl,Duże rozmiary butów,tęgie, szerokie, tęgość H,duże buty 41-46 ,na zamówienie, tęgie buty, Buty damskie duże rozmiary,buty na tęgą stopę,tęgość H,na tęgie stopy,na szeroką stopę,na szerokie stopy,na dużą stopę,na duże stopy,szerokie,tęgie,płaskie

Buty xxl,Duże rozmiary butów,tęgie, szerokie, tęgość H,duże buty 41-46 ,na zamówienie, tęgie buty, Buty damskie duże rozmiary,buty na tęgą stopę,tęgość H,na tęgie stopy,na szeroką stopę,na szerokie stopy,na dużą stopę,na duże stopy,szerokie,tęgie,płaskie

DUŻE BUTY DAMSKIE VICTORIA SHOES 41 42 43 44 45 46 CZÓŁENKA BEŻOWO CZARNE . www.victoriashoes.pl na miarę i na zamówienie | victoriashoes.pl

Victoria shoes buty damskie - https://pegazimielski.pl/ A – buty damskie duże rozmiary sklep , obuwie damskie rozmiar 42 43 44 45 , buty damskie na zamówienie , buty xxl damskie , duże
Producent obuwia ROBERTO - sklep internetowy, DUŻE BUTY, buty damskie, obuwie męskie, buty młodzieżowe, wielkie buty, nietypowe rozmiary obuwia, duże rozmiary butów, wielkie rozmiary,duże obuwie, rozmiar xl, rozmiary xxl, dla wielkich, e-sklep,

Duże rozmiary butów damskich ,szerokie tęgie nietypowe buty XXL, na szeroką stopę,duże haluksy ,na tęgie opuchnięte chore stopy , tęgość H,K ,na zamówienie rozmiar 42,43,44,45,46, od szewca,botki,kozaki, czółenka

Deloito damskie duże rozmiary klamra pasek na kostkę sandały kliny letnie tkactwo jednolity kolor oddychające buty, - Gorący róż - 38 EU : Amazon.pl: Moda

Europejska i amerykańska moda i wygodne jednoetapowe buty damskie Duże rozmiary butów na grubym obcasie kup niedrogo — cena, bezpłatna dostawa, autentyczne opinie ze zdjęciami — Joom

2022 nowych moda zima nowe buty za kostkę damskie duże rozmiary gąbka dół kobiet buty okrągłe Toe sznurowane buty damskie|Buty do kostki| - AliExpress