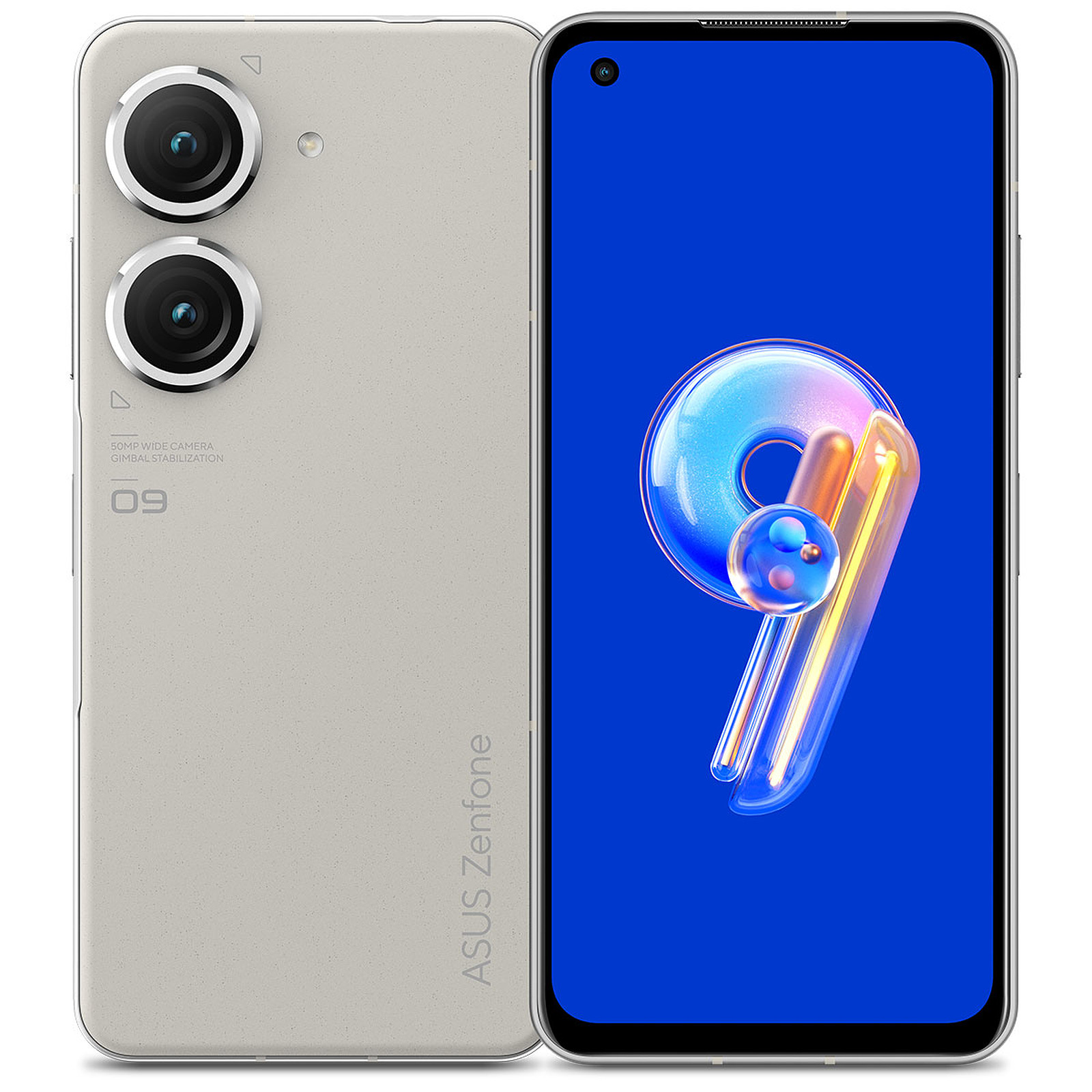
Amazon.com: Asus Zenfone 7 Pro (ZS671KS) 5G 256GB 8GB Global Edition - Aurora Black : Cell Phones & Accessories

HUANGTAOLI Cover per ASUS Zenfone 7 ZS670KS, Cover a Libro con Funzione Stand e Porta Carte Chiusura Magnetica Portafoglio (6.67 Pollici) : Amazon.it: Elettronica

Asus Fonepad FE170CG-1B012A - Tablet (17,7 cm/7 Pulgadas, Intel Atom Z2520, 1,2 GHz, 1GB RAM, Memoria Interna de 8 GB, Android 4.3) Blanco Blanco : Amazon.es: Informática

Amazon.com: Rantow Foldable Phone/Tablet Monitor Sunshade for DJI Mavic Pro/Mavic Air/Mavic 2 Zoom/Mavic 2 Pro/Spark/Phantom 3 4 / Inspire 1/OSMO Accessories Controller Sun Hood : Cell Phones & Accessories

LCD originale da 5.7 pollici per ASUS Zenfone AR ZS571KL Display LCD Touch Screen Digitizer Assembly con cornice per ASUS ZS571KL LCD|Schermi LCD per cellulare| - AliExpress

Asus ME173X-1F010A MeMO Pad HD, Pannello LCD da 7 Pollici LED IPS, Processore Quad Core 1.2GHz, 16GB di SSD , Verde | seguiprezzo/segui Amazon , grafici cronologia prezzo Amazon, monitoraggi prezzo Amazon,

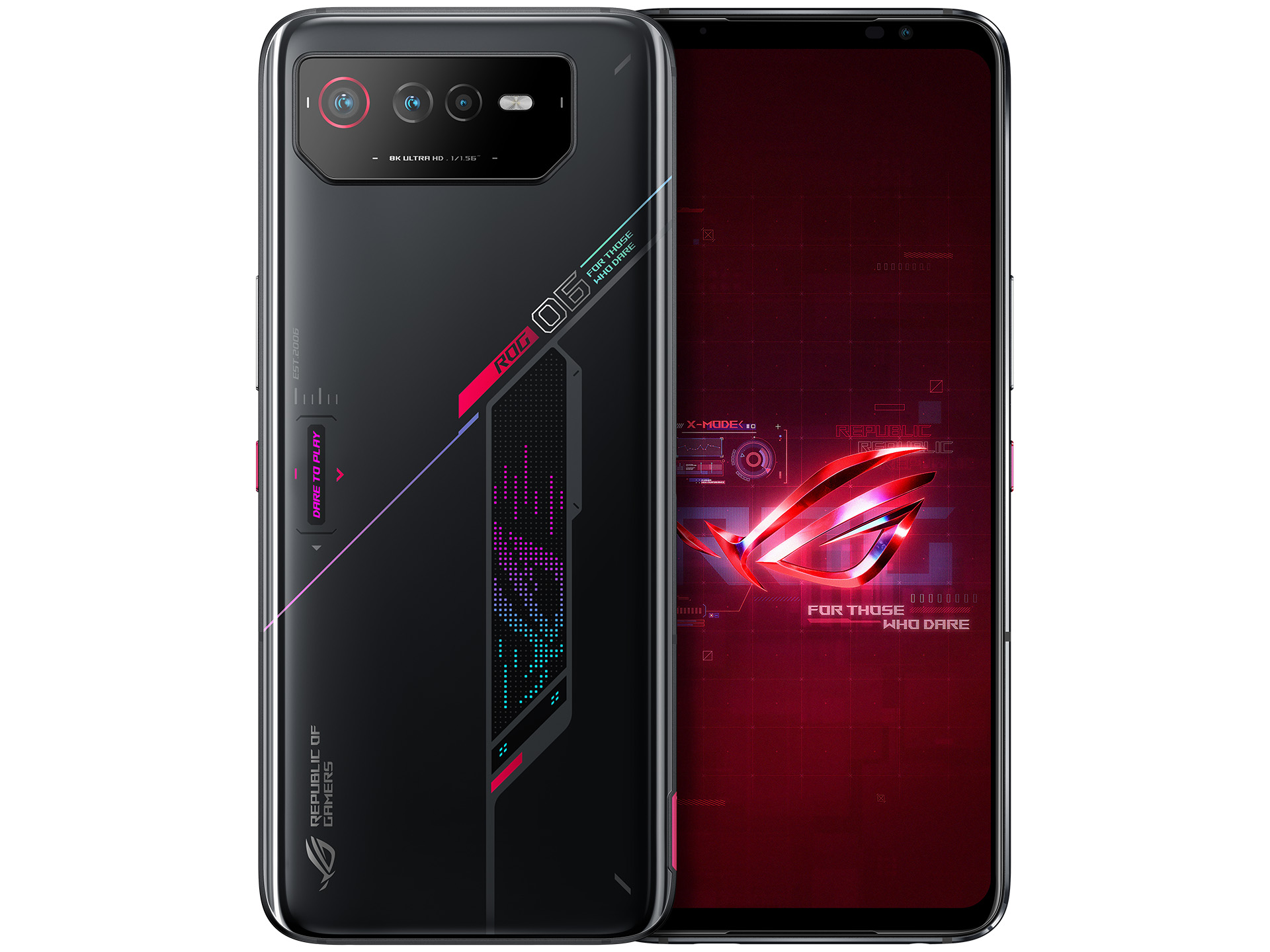
Amazon.com: ASUS ROG Phone 7 5G Dual SIM 512GB 16GB RAM Factory Unlocked (GSM Only | No CDMA - not Compatible with Verizon/Sprint) Global Version - White : Cell Phones & Accessories

ASUS Zenfone 8 Compact 5G Smartphone (15 cm (5,9 pollici) Full HD+ 120 Hz AMOLED Display, fotocamera da 64 MP, 8 GB di RAM, 256 GB di memoria) Nero : Amazon.it: Elettronica

Originale 7 ''pollici per Asus MeMO Pad HD7 ME173 ME173X K00B pannello Touch Screen Digitizer sostituzione lente in vetro nero|Schermi LCD e pannelli per tablet| - AliExpress

Amazon.com: ASUS ROG Phone 7 5G Dual SIM 512GB 16GB RAM Factory Unlocked (GSM Only | No CDMA - not Compatible with Verizon/Sprint) Global Version - White : Cell Phones & Accessories







![Custodia Smart Folio per iPad Air 10.9 per iPad Air 4°, 5° / iPad Pro 1° [3 colori] – elago Custodia Smart Folio per iPad Air 10.9 per iPad Air 4°, 5° / iPad Pro 1° [3 colori] – elago](https://cdn.shopify.com/s/files/1/0498/9589/9290/products/EPADP129-5-MFLO-SPK_900x.jpg?v=1675452672)


