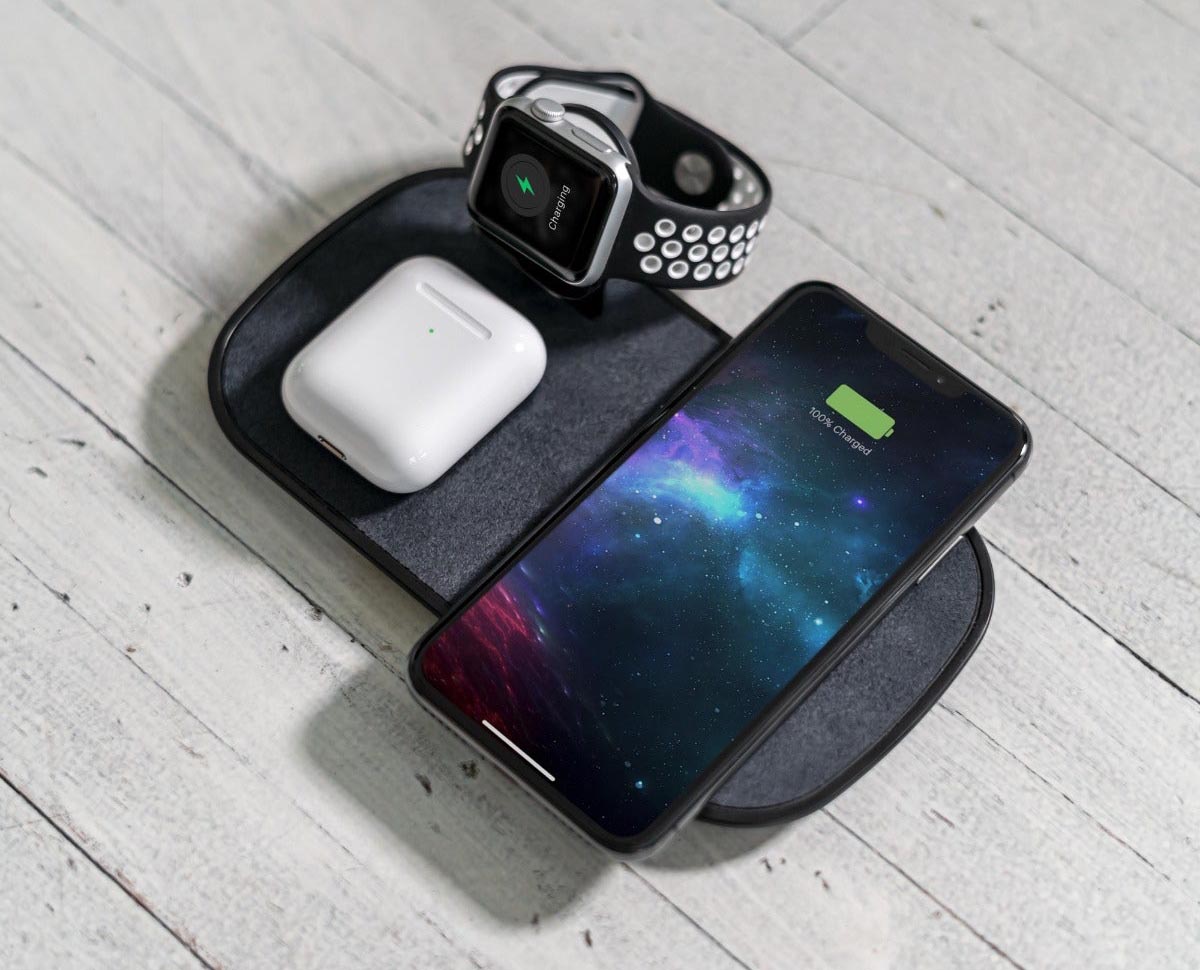
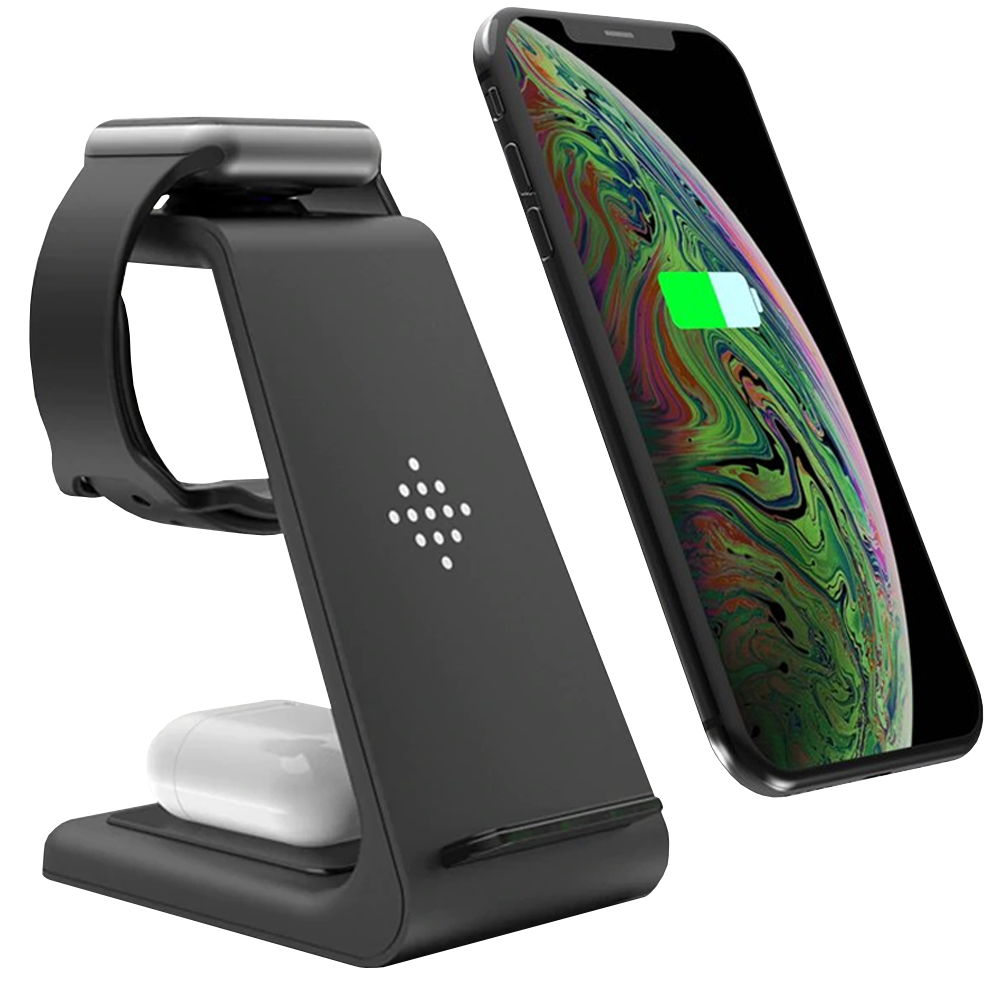
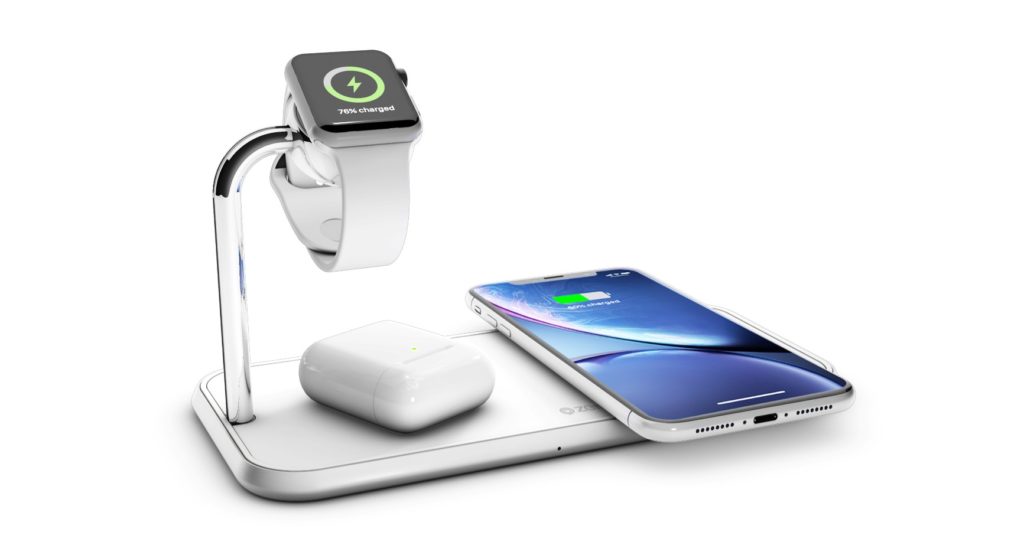
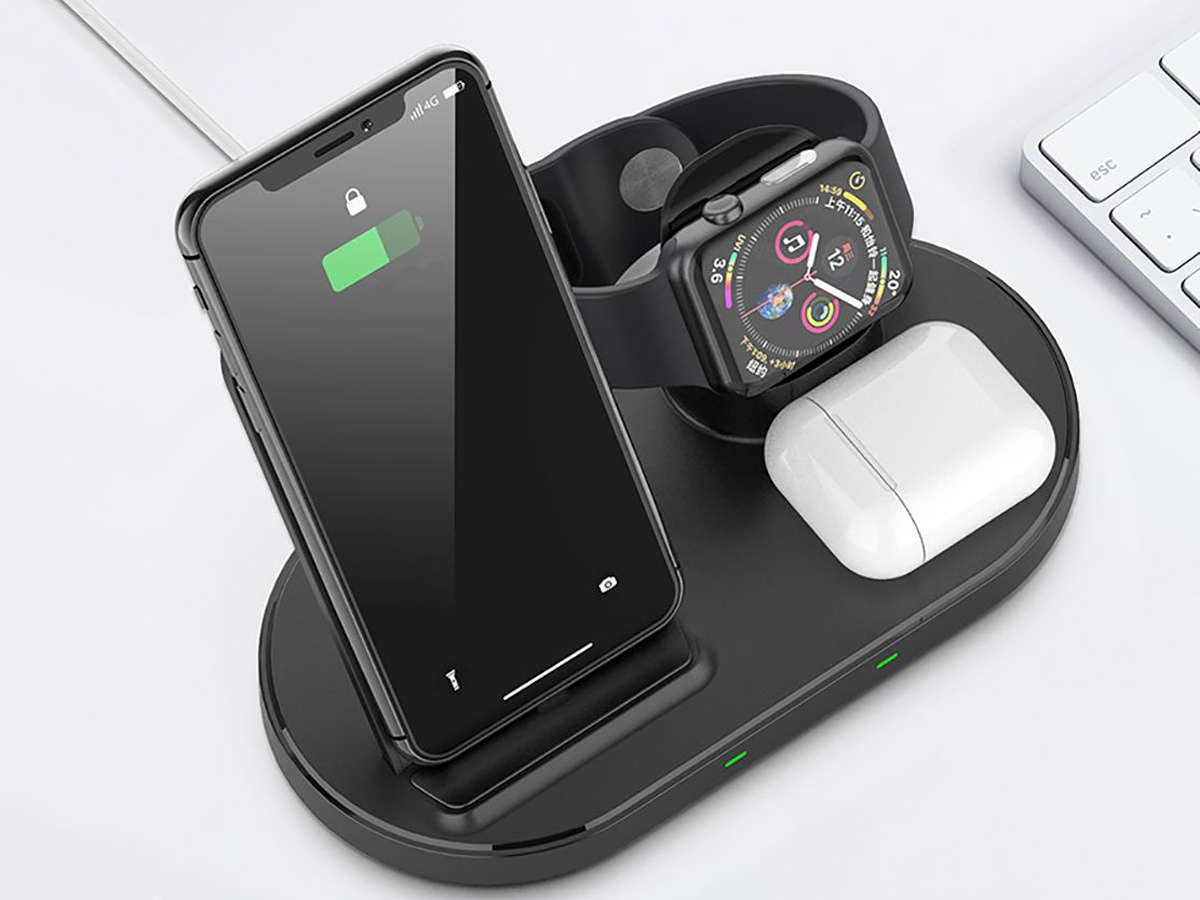
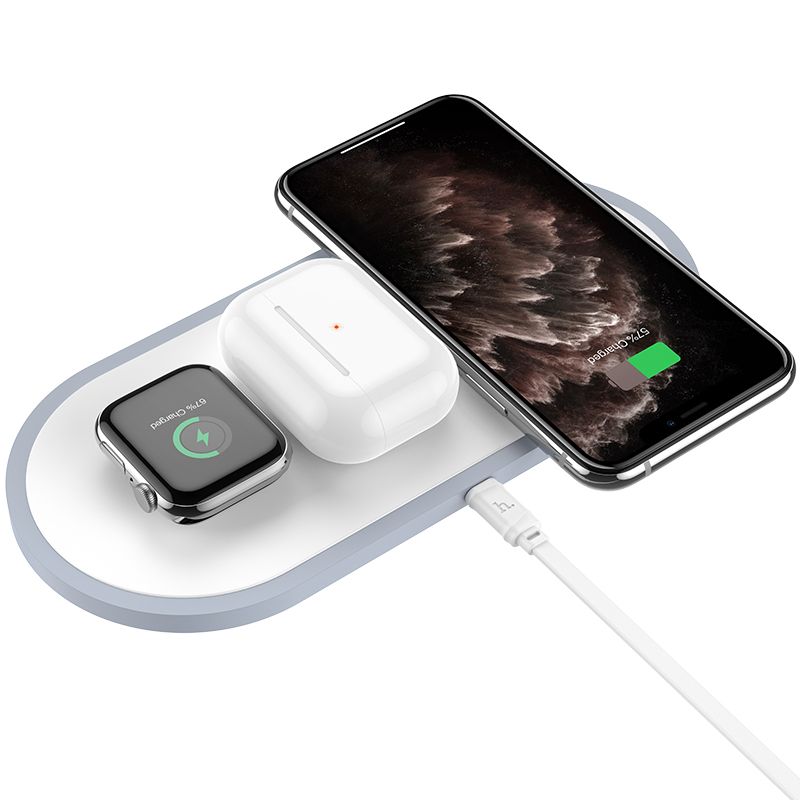
Recenzja Wireless Charge Dock 2 in 1 z GearBest - ładowarka indukcyjna dla iPhone i Apple Watch bez użycia kabli z zestawu - Serwis iPhone Szczecin - AppleMobile.pl

Ładowarka indukcyjna Belkin Boost MFi 3-in-1 Qi 15W dla Apple Watch / AirPods / iPhone z MagSafe, czarna Pancernik.eu 745883819461

Belkin Ładowarka indukcyjna (iPhone, Apple Watch, biała) - Ładowarki do smartfonów - Sklep komputerowy - x-kom.pl
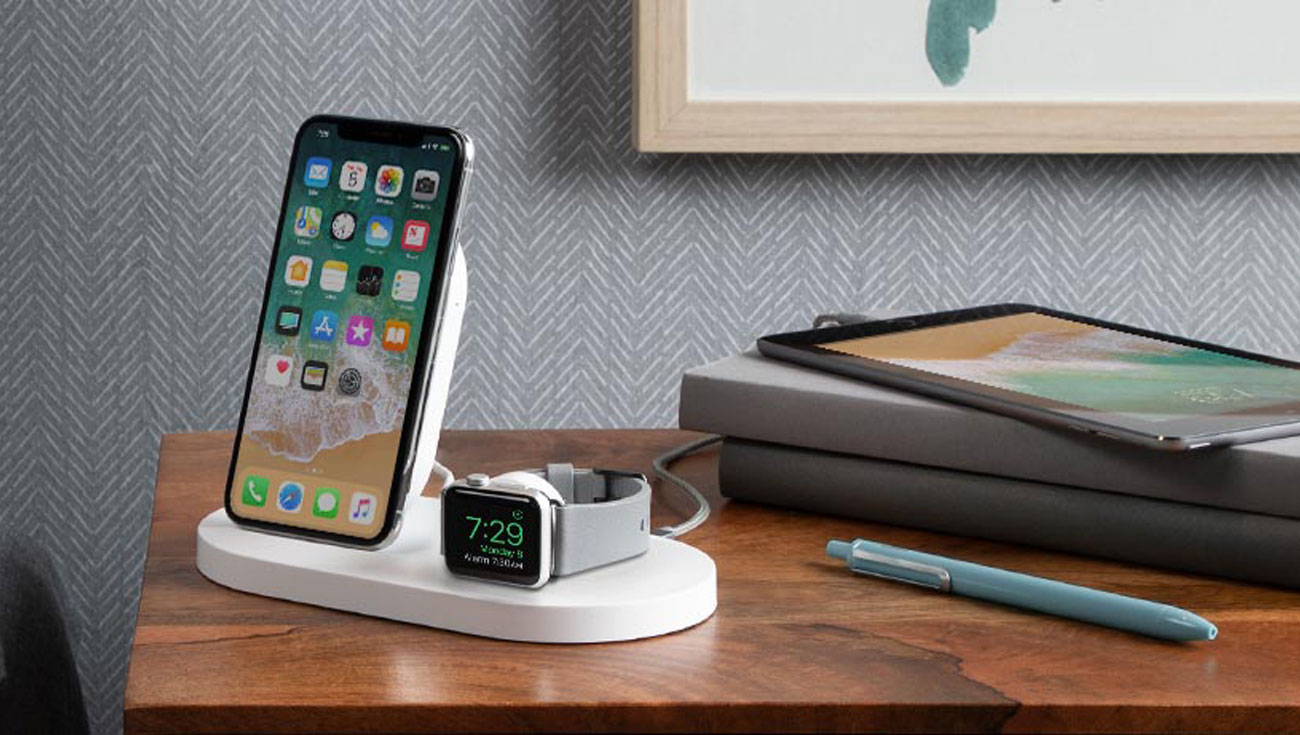

Belkin Ładowarka indukcyjna (iPhone, Apple Watch, czarna) - Ładowarki do smartfonów - Sklep komputerowy - x-kom.pl
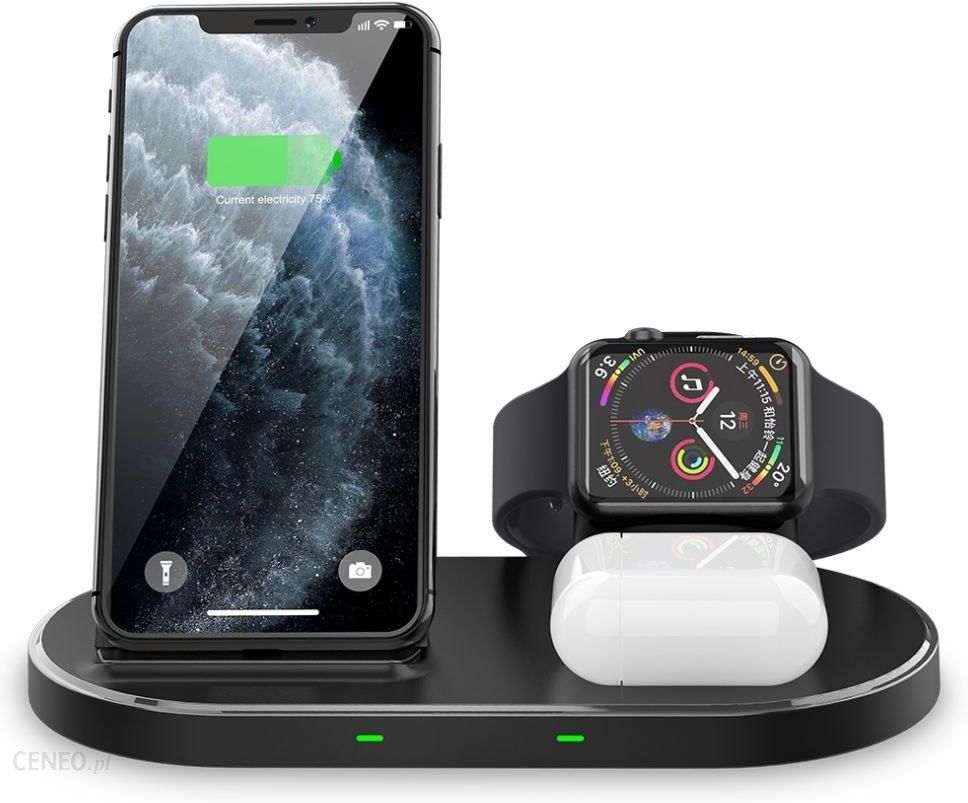

Ładowarka do telefonu Tech-Protect Ładowarka Indukcyjna W55 Do Iphone'A Lub Androida Airpodsa I Apple Watch - Opinie i ceny na Ceneo.pl
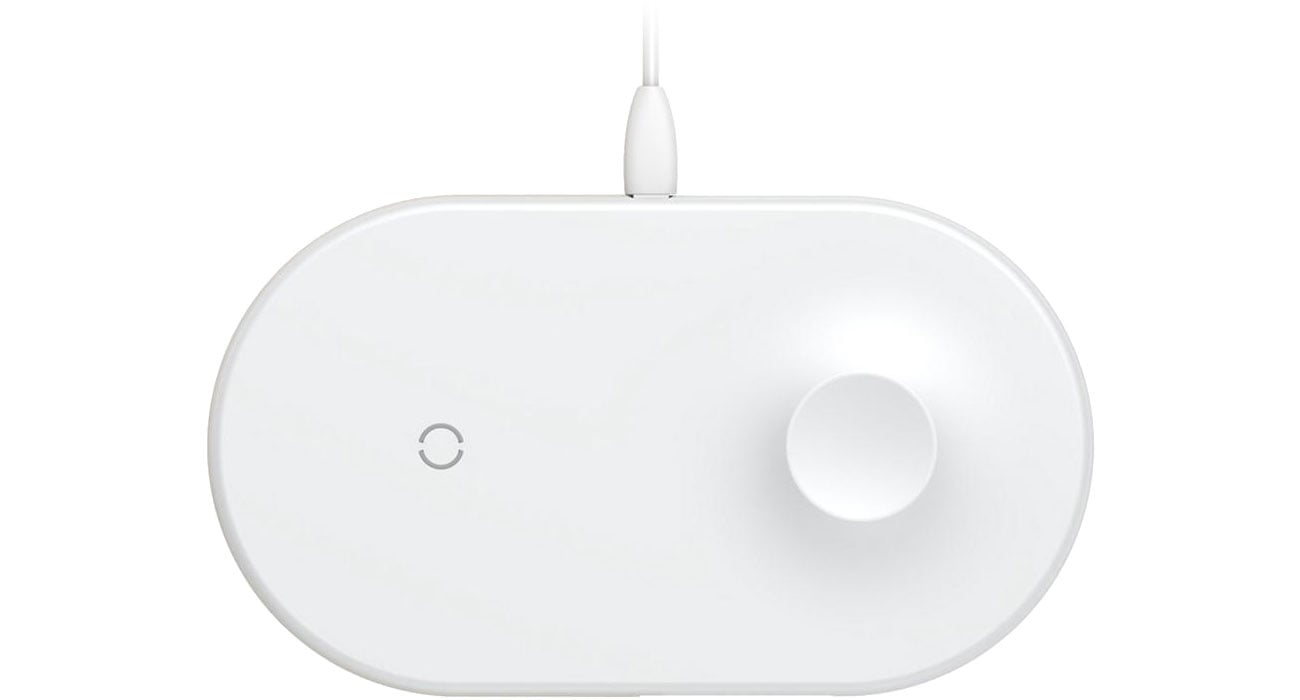

Baseus Ładowarka indukcyjna 10W (Smartphone+Apple Watch) - Ładowarki do smartfonów - Sklep komputerowy - x-kom.pl
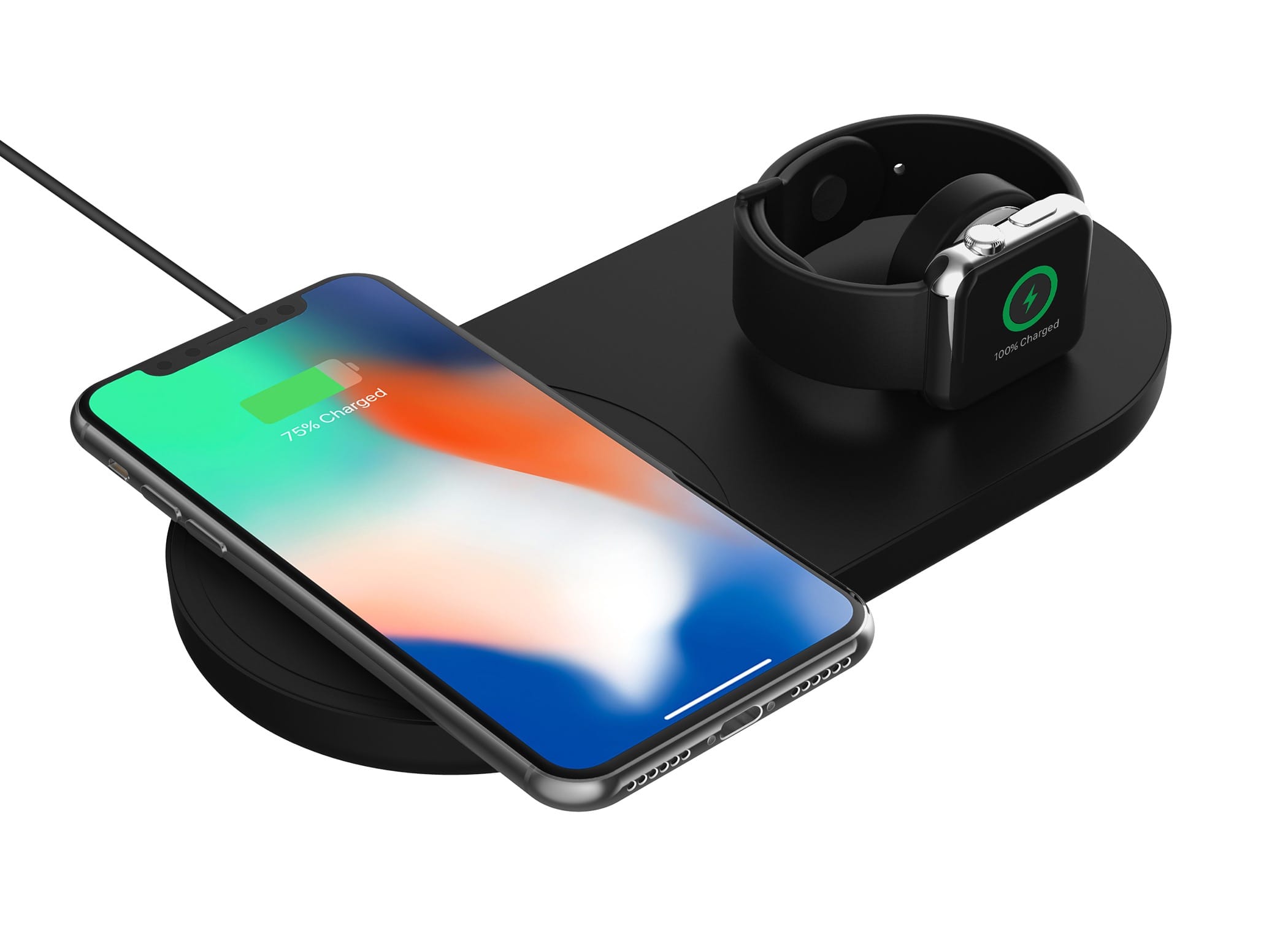

Nowe akcesoria Griffin – powerbank, ładowarki Qi dla iPhone i Apple Watch oraz kable USB-C do Lightning | iMagazine

Bezprzewodowa ładowarka Belkin BOOST UP CHARGE 3-in-1 Wireless Charger do iPhone'a, Apple Watch i AirPods – czarna - Apple (PL)

Baseus Smart 3in1 / Ładowarka indukcyjna bezprzewodowa Qi do iPhone Apple Watch AirPods 3w1 18W Max EOL | Zasilanie \ Ładowarki indukcyjne Nieprzecenione Wybrane ładowarki -20% Nieprzecenione -10%
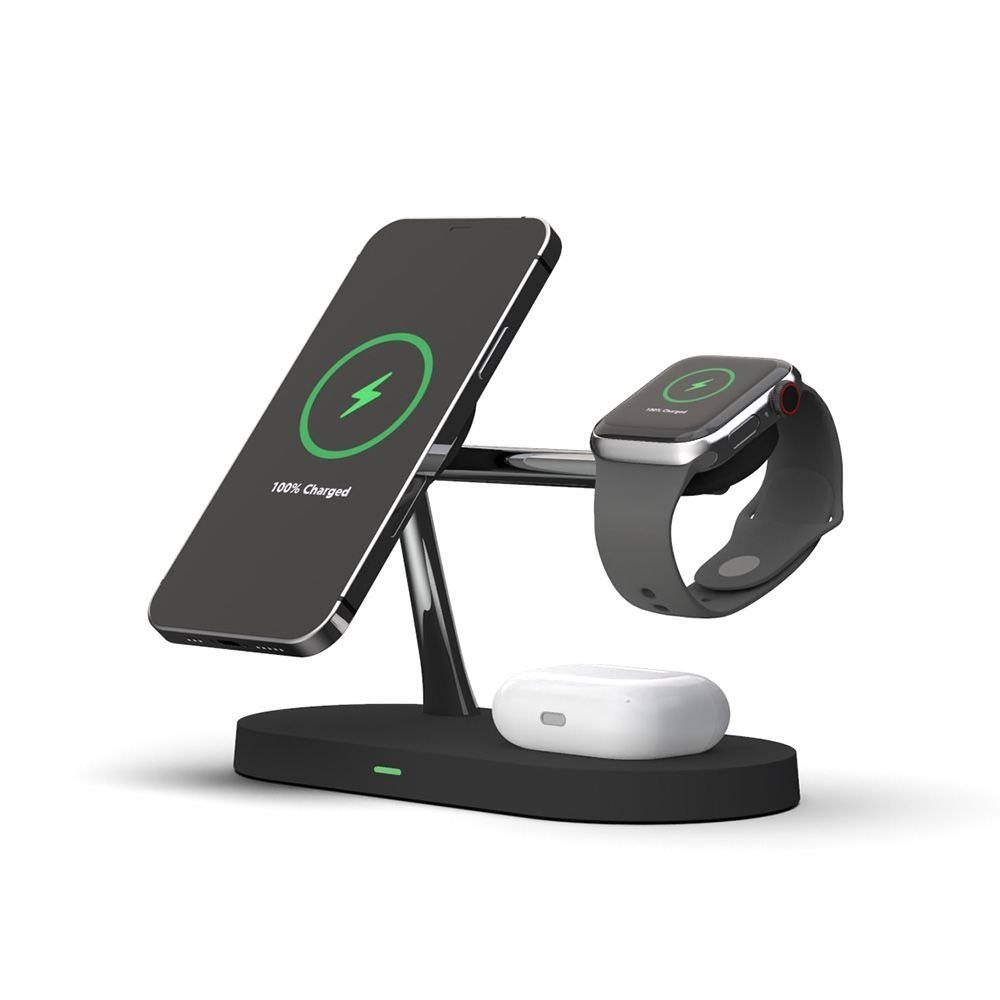

Wielofunkcyjna ładowarka indukcyjna 3w1 Magsafe, Airpods, Apple Watch z lampką LED - TECH-PROTECT | Sklep EMPIK.COM