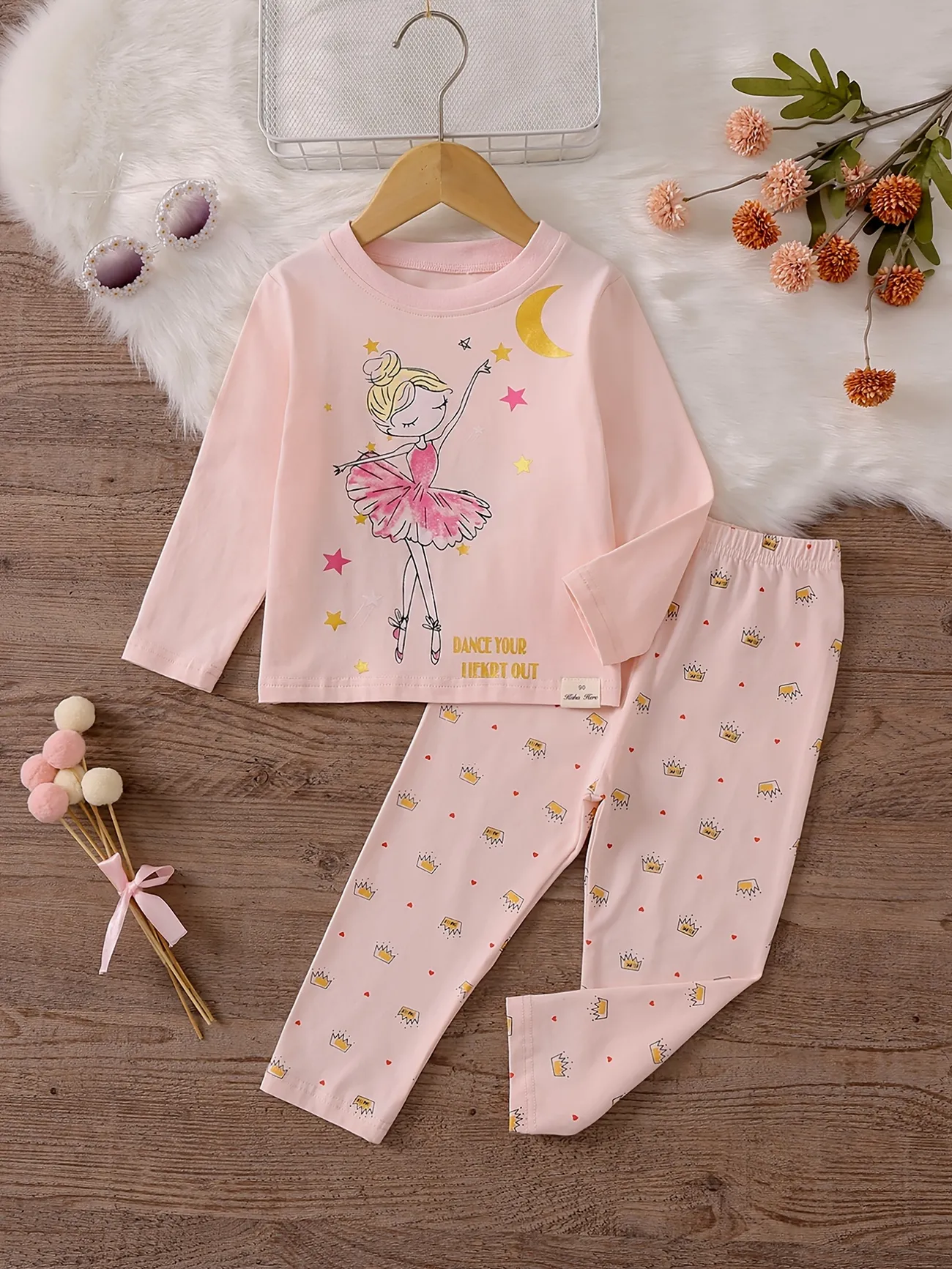
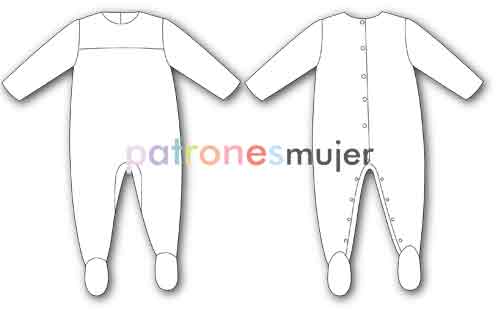
Amazon.com: AVAUMA - Conjunto de pijama para bebés y niñas 6M-8T para niños lindos para niños pequeños ajustados Pjs algodón ropa de dormir : Ropa, Zapatos y Joyería
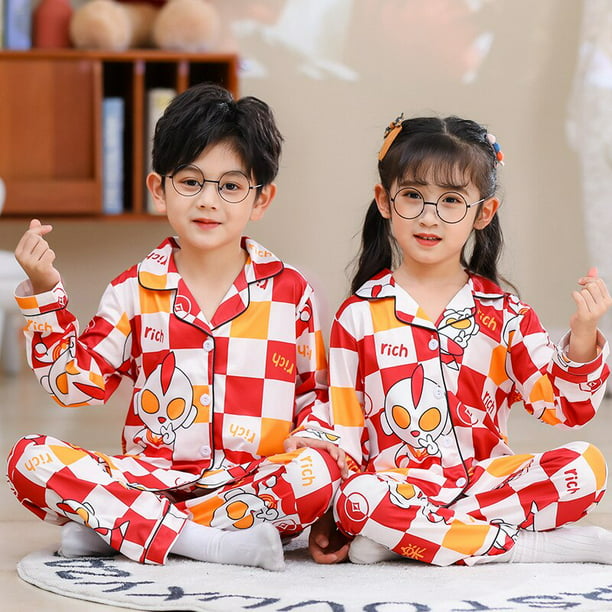

Disney Toy Story Three Eyes Conjuntos de pijamas de satén para niños Pijamas para niños Ropa de dormir para bebés Pijamas para niñas Pijamas para adolescentes Pijamas para niñas PjsS altura 145-155cm

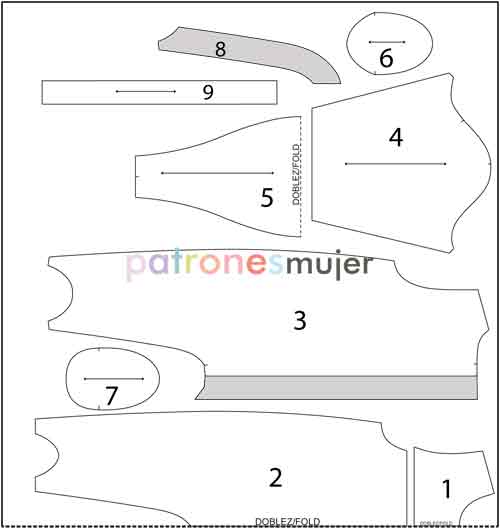
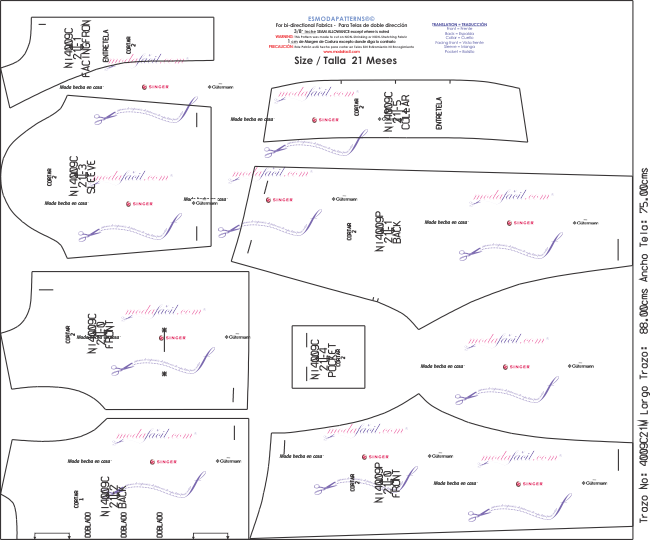
Amazon.com: Simplicity Crafts 5276 - Pijama de muñeca de bebé, patrón de costura para niñas, de Andrea Schewe, talla 18 pulgadas : Arte y Manualidades