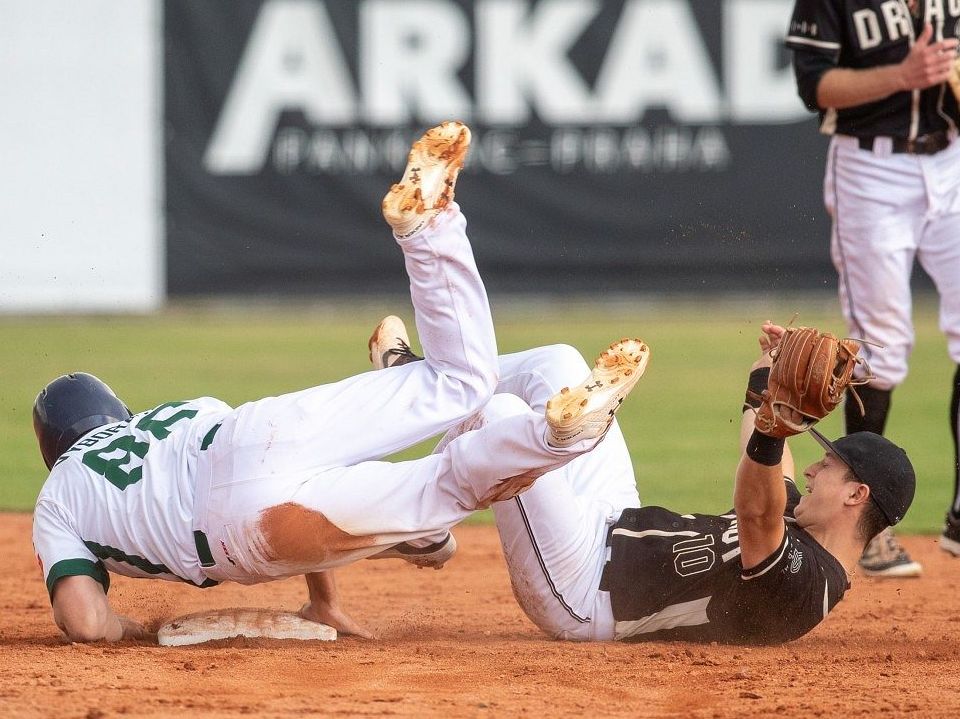
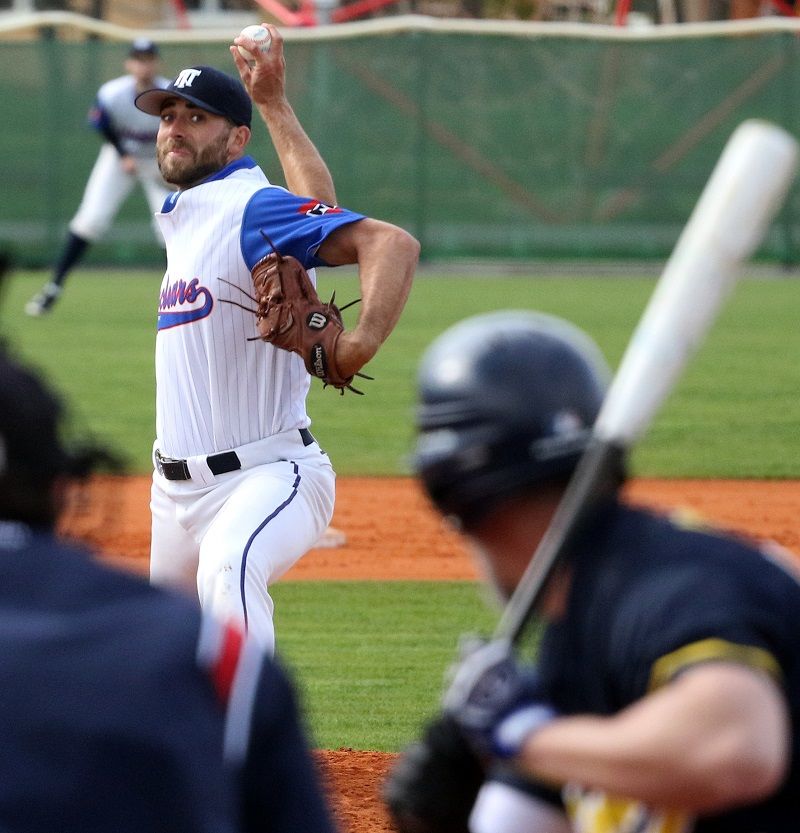
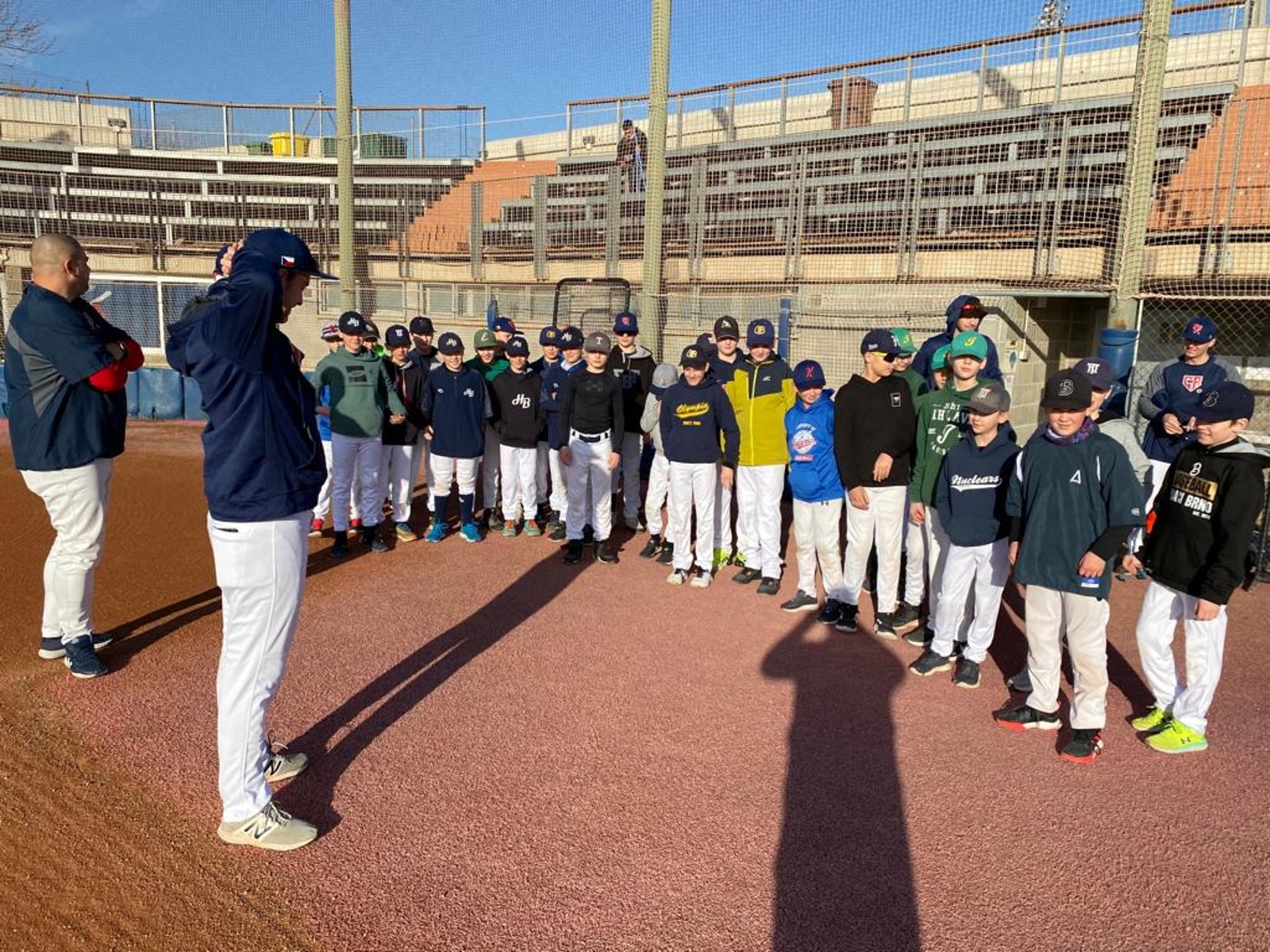
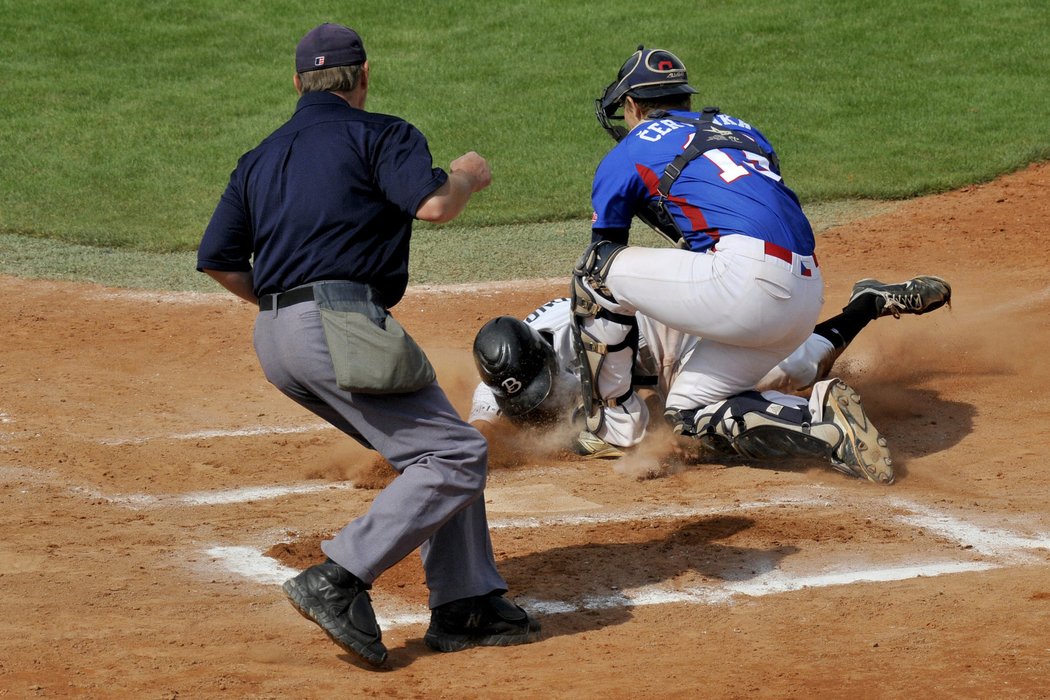
Druhou sérii baseballové Extraligy prohrál Sokol Hluboká s úřadujícím mistrem Draci Brno | Sport | Budějcká Drbna - zprávy z Českých Budějovic a jižních Čech
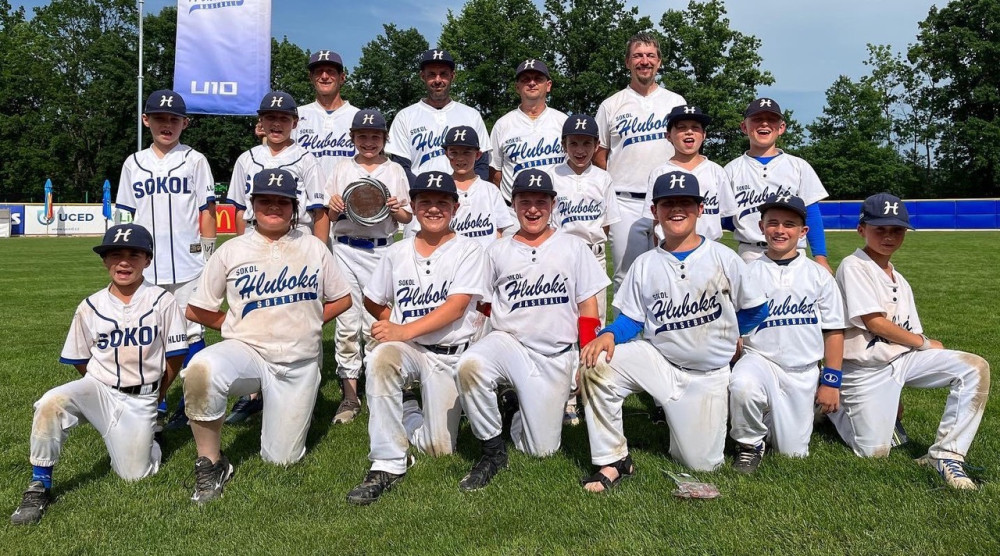

Hlubocké baseballové léto budiž pochváleno | Z kraje | Budějcká Drbna - zprávy z Českých Budějovic a jižních Čech
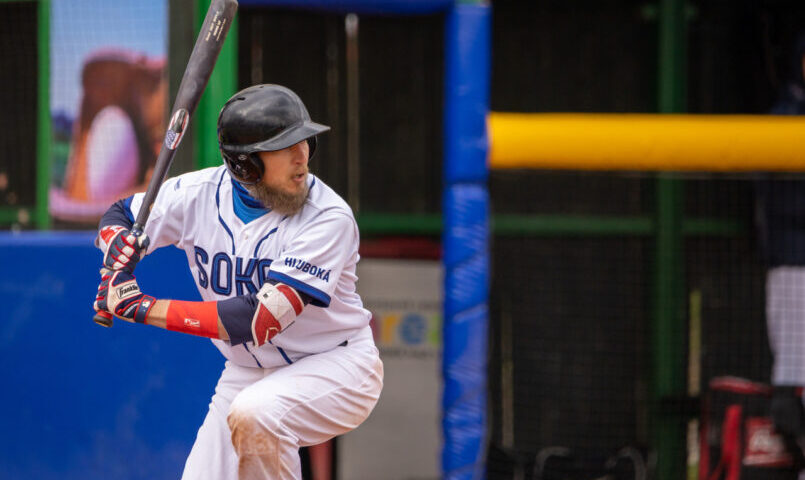

Na Hlubokou přiletí brněnští Draci, úřadující mistři a největší favorit baseballové extraligy! | Milujeme Baseball
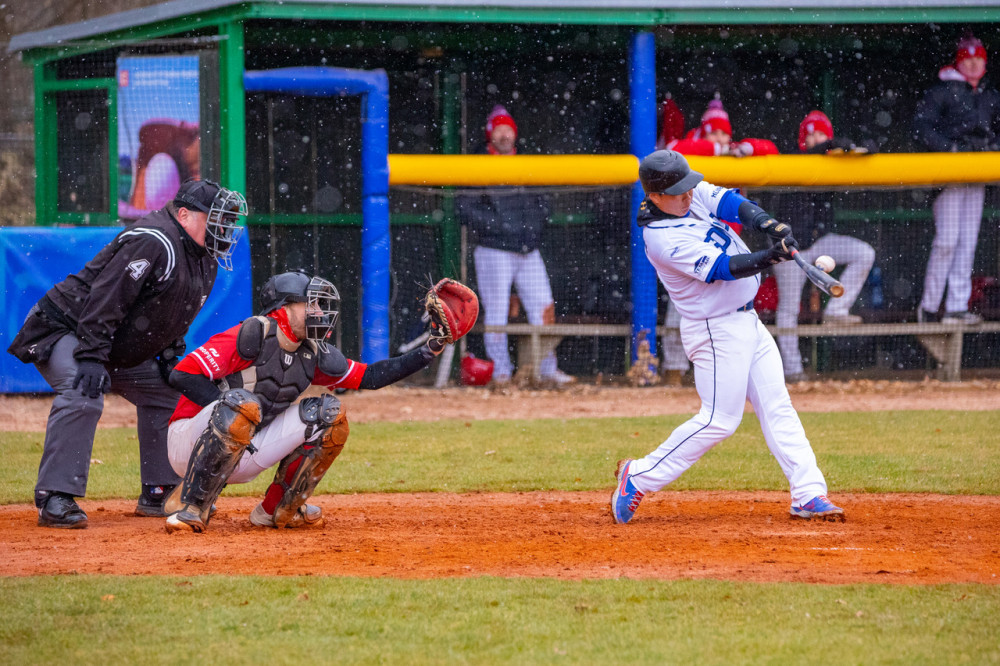

Sokol Hluboká vletěl do baseballové Extraligy a za sněžení porazil doma Techniku Brno | Ostatní sporty | Sport | Budějcká Drbna - zprávy z Českých Budějovic a jižních Čech
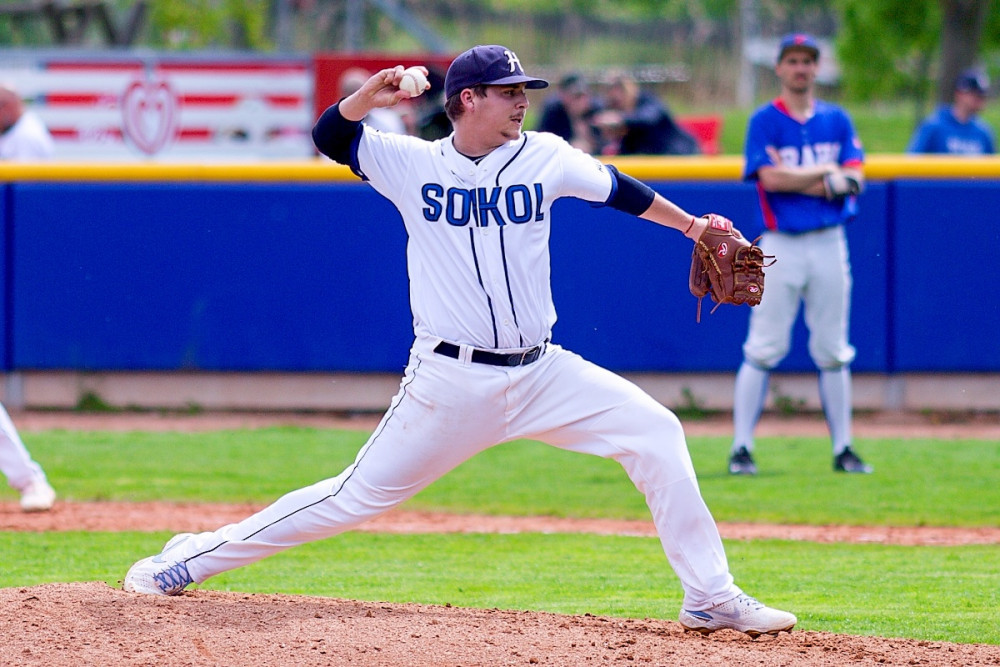

Dohrávka čtvrtého kola mezi Jabloncem nad Nisou a Hlubokou byla opět odložena, v sobotu přivítá Sokol brněnské Hrochy | Ostatní sporty | Sport | Budějcká Drbna - zprávy z Českých Budějovic a jižních Čech
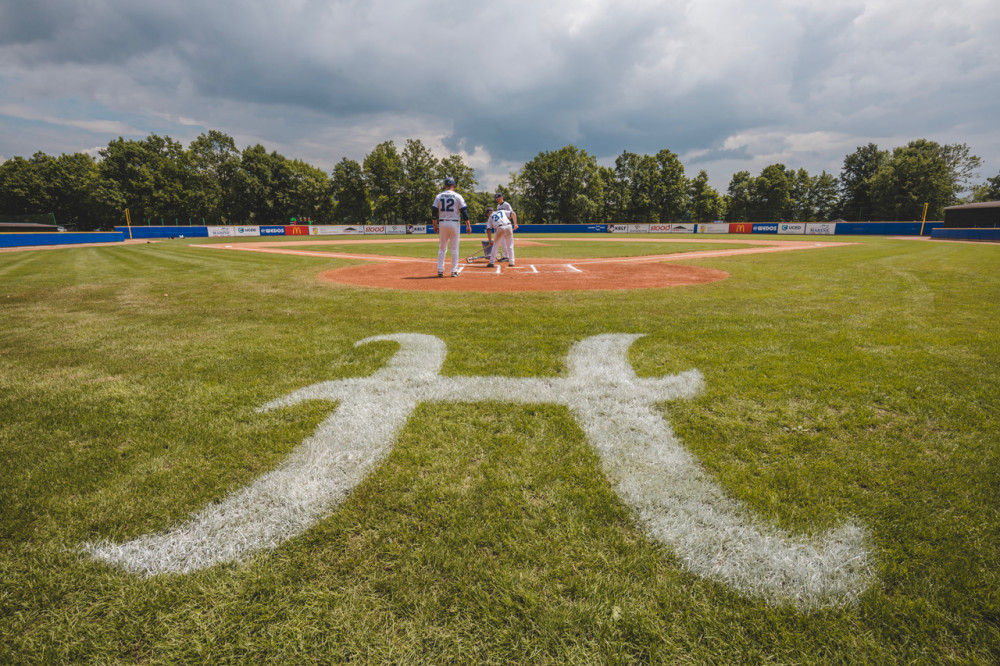

Druhou sérii baseballové Extraligy prohrál Sokol Hluboká s úřadujícím mistrem Draci Brno | Sport | Budějcká Drbna - zprávy z Českých Budějovic a jižních Čech