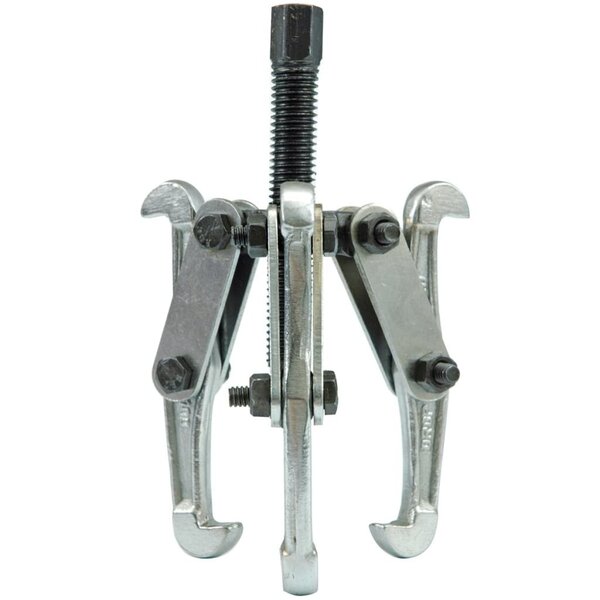
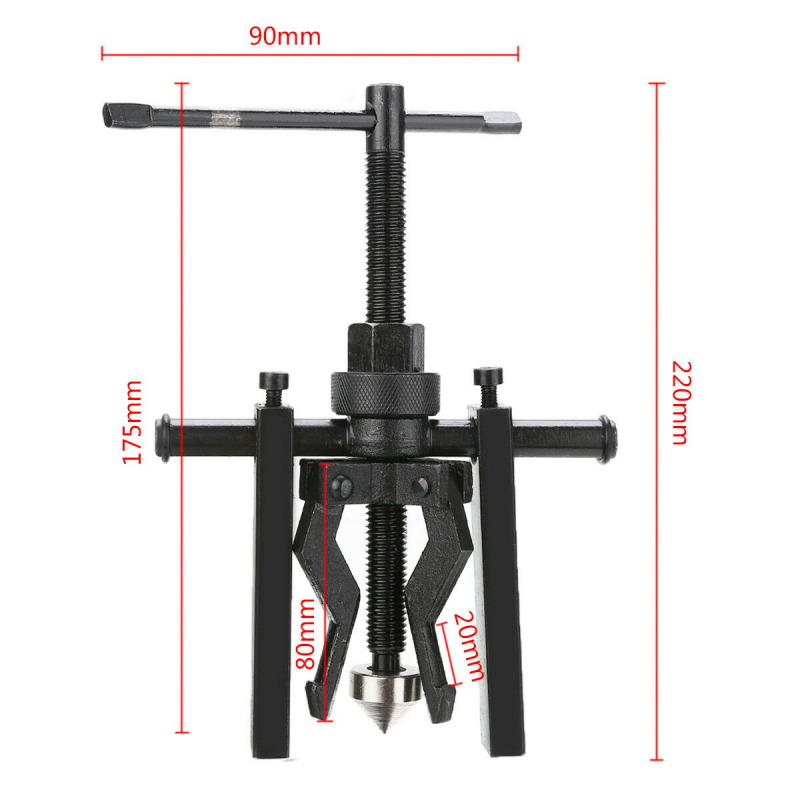
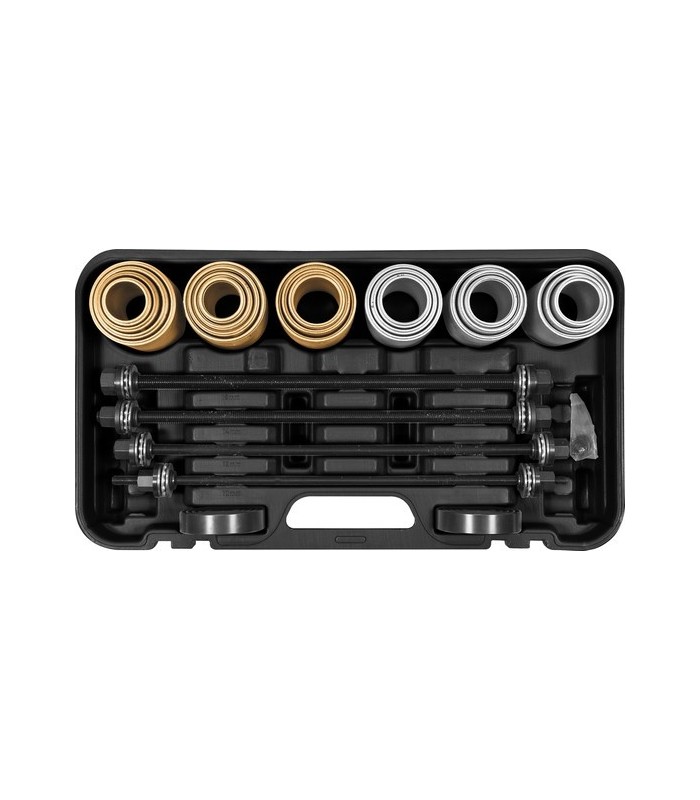
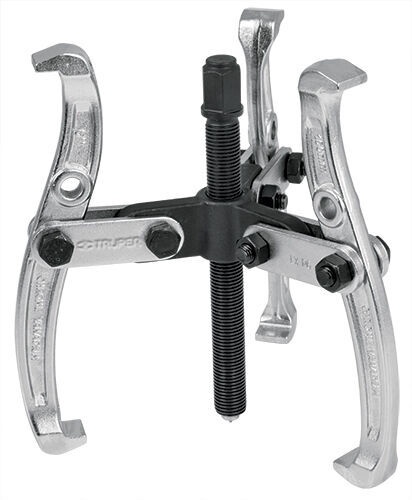
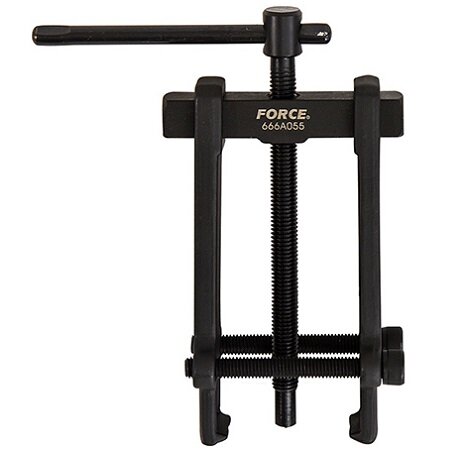
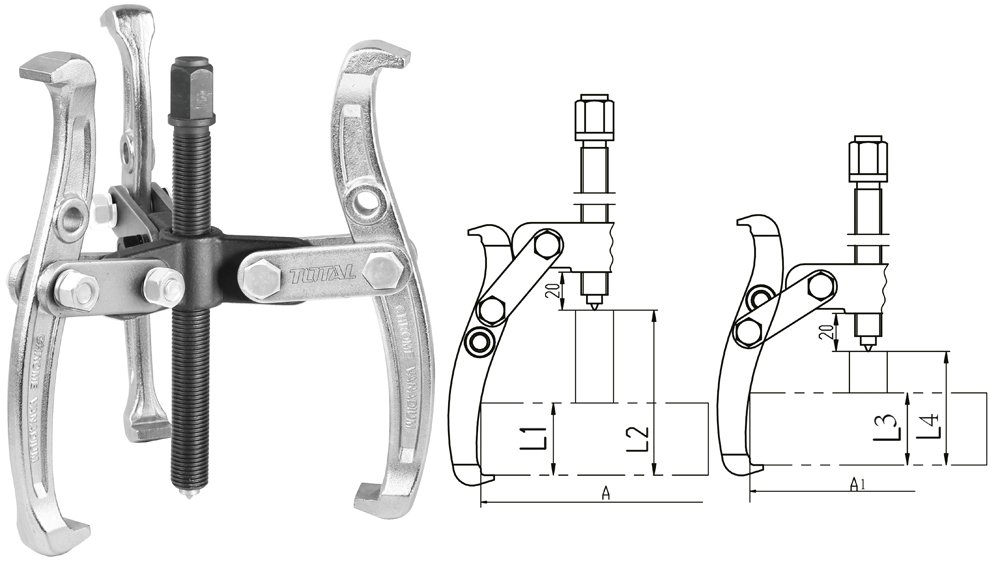
Guolių nuėmėjas su hidrauliniu cilindru 5t. | Nuėmėjų rinkiniai | Guolių, skremulių nuėmėjai. | Įrankiai auto remontui | Irankiu Baze
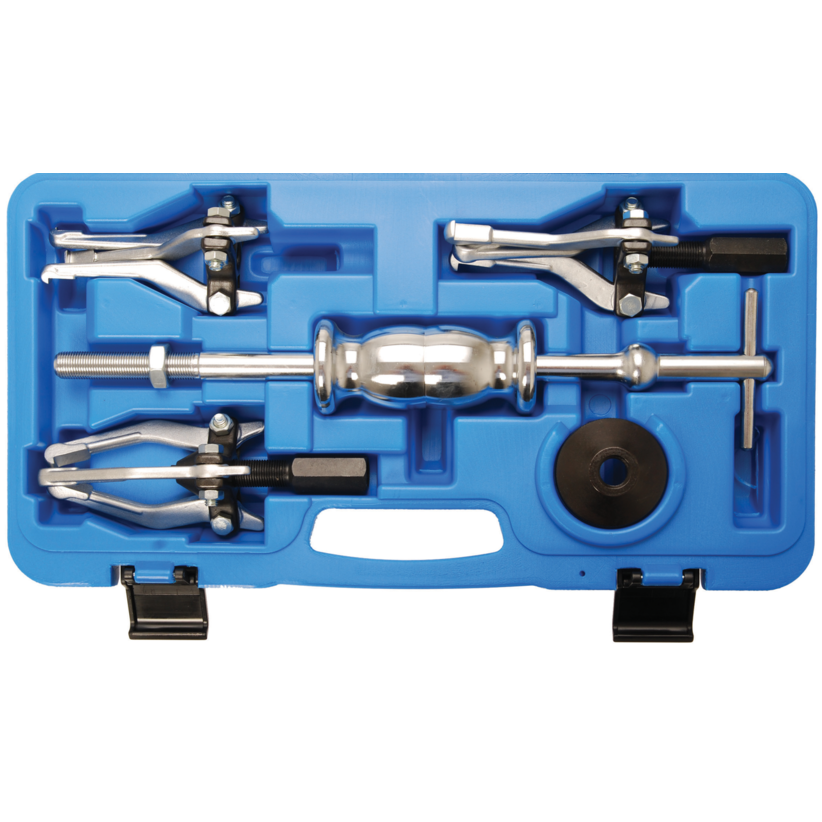

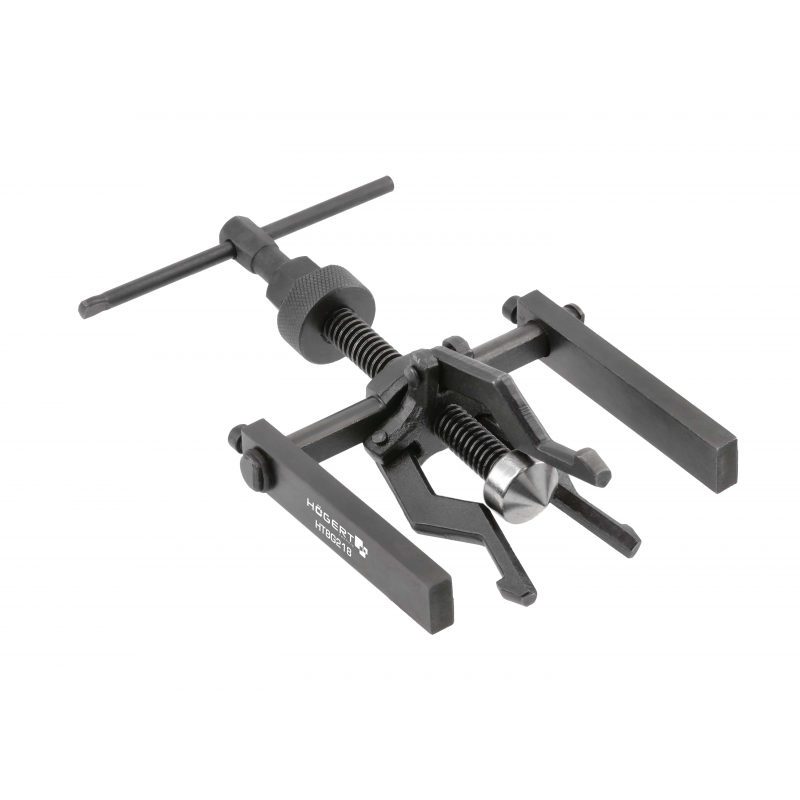
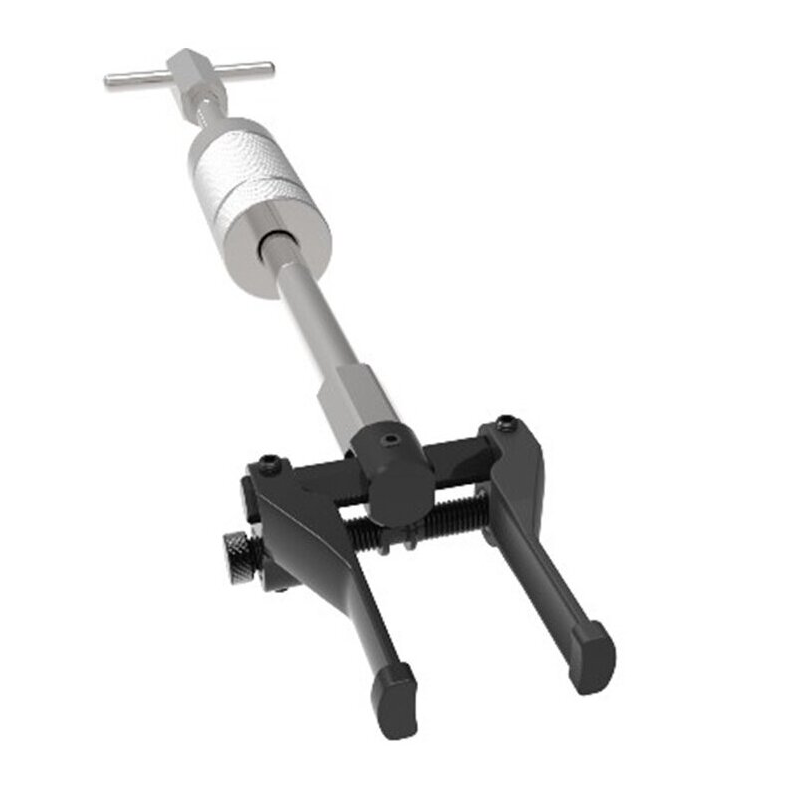
Vidinių, išorinių guolių nuėmėjas su atbuliniu plaktuku | Nuėmėjų rinkiniai | Guolių, skremulių nuėmėjai. | Įrankiai auto remontui | Irankiu Baze