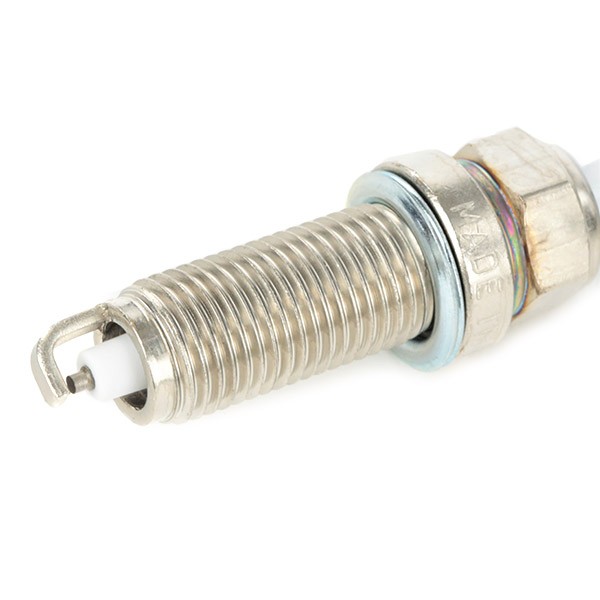
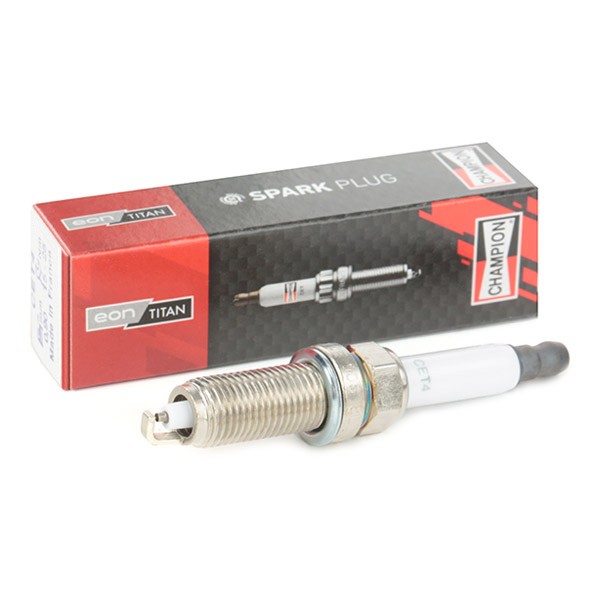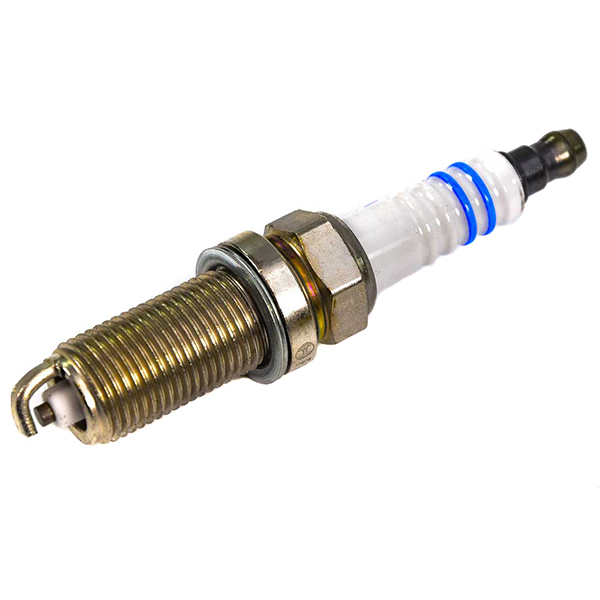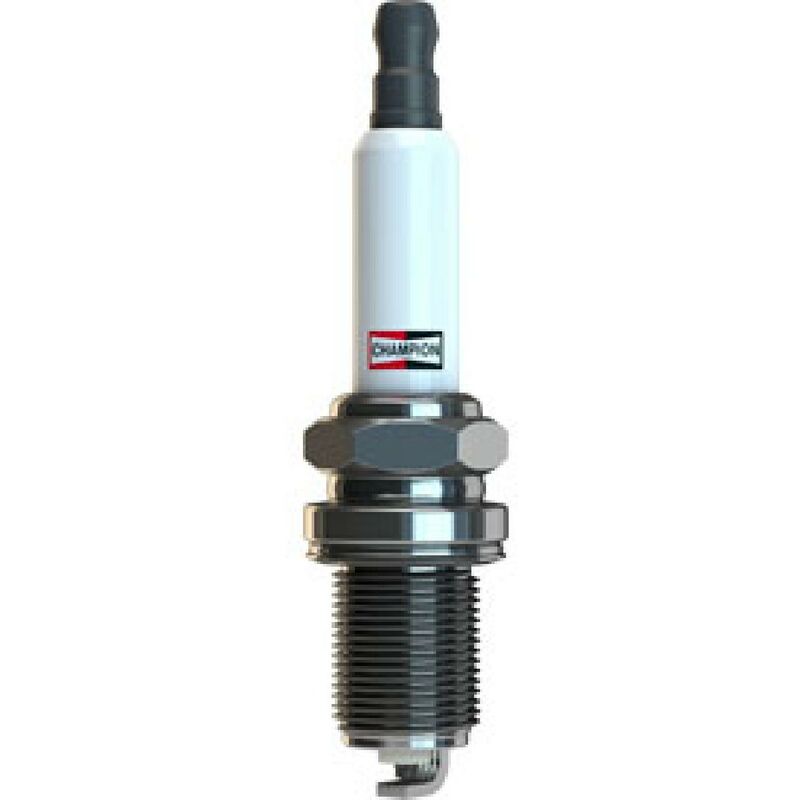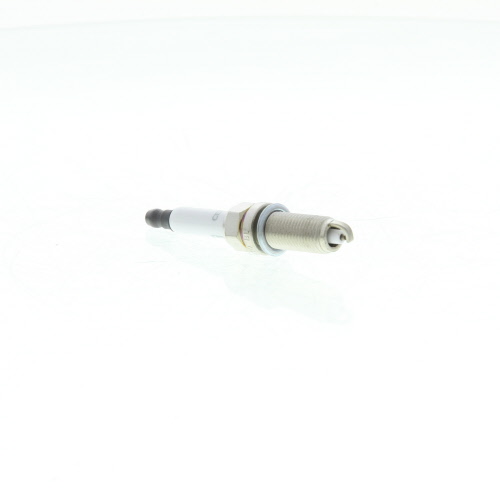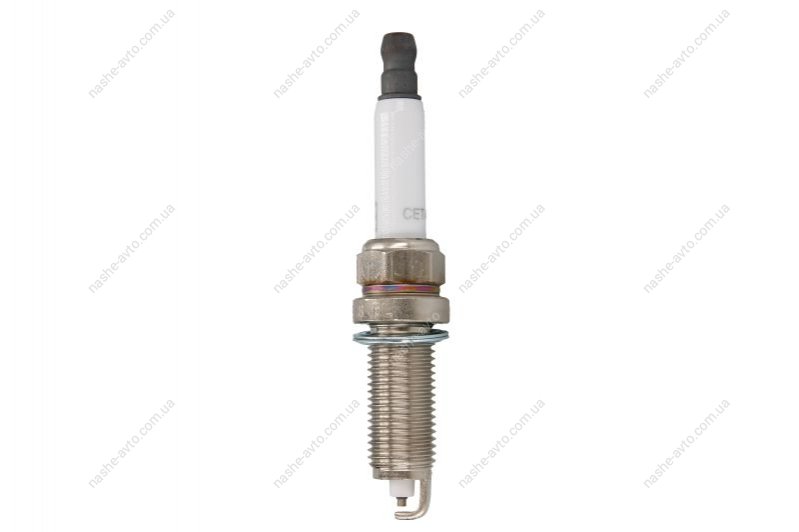
CET4 CHAMPION Aerovantage Spoiler Spark plug CET4, M12x1.25, Spanner Size: 14 mm, Ti Poly-V ▷ AUTODOC price and review

Sofascore on X: "🗓️ | SCHEDULE Finals, finals and more finals — that's what's coming up today and tomorrow in the world of football! 🤩 There are national cup finals in England,

IM24CA World Council - Annual General Meeting 2023 - summary and voting results | International Melges 24 Class Association
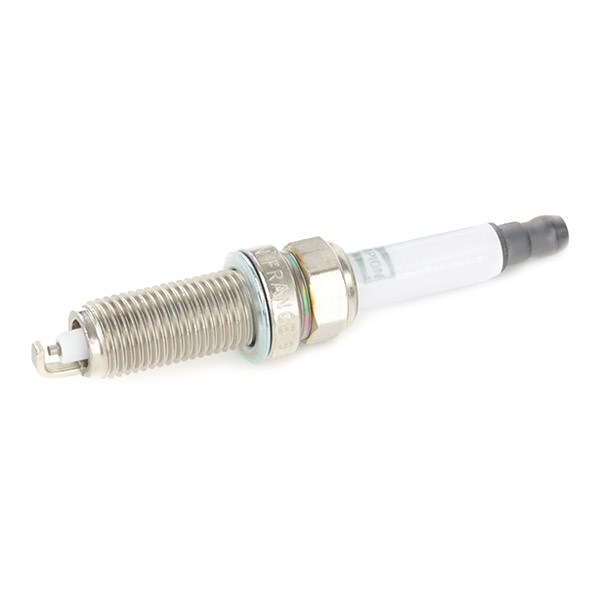

CET4 CHAMPION Aerovantage Spoiler Spark plug CET4, M12x1.25, Spanner Size: 14 mm, Ti Poly-V ▷ AUTODOC price and review
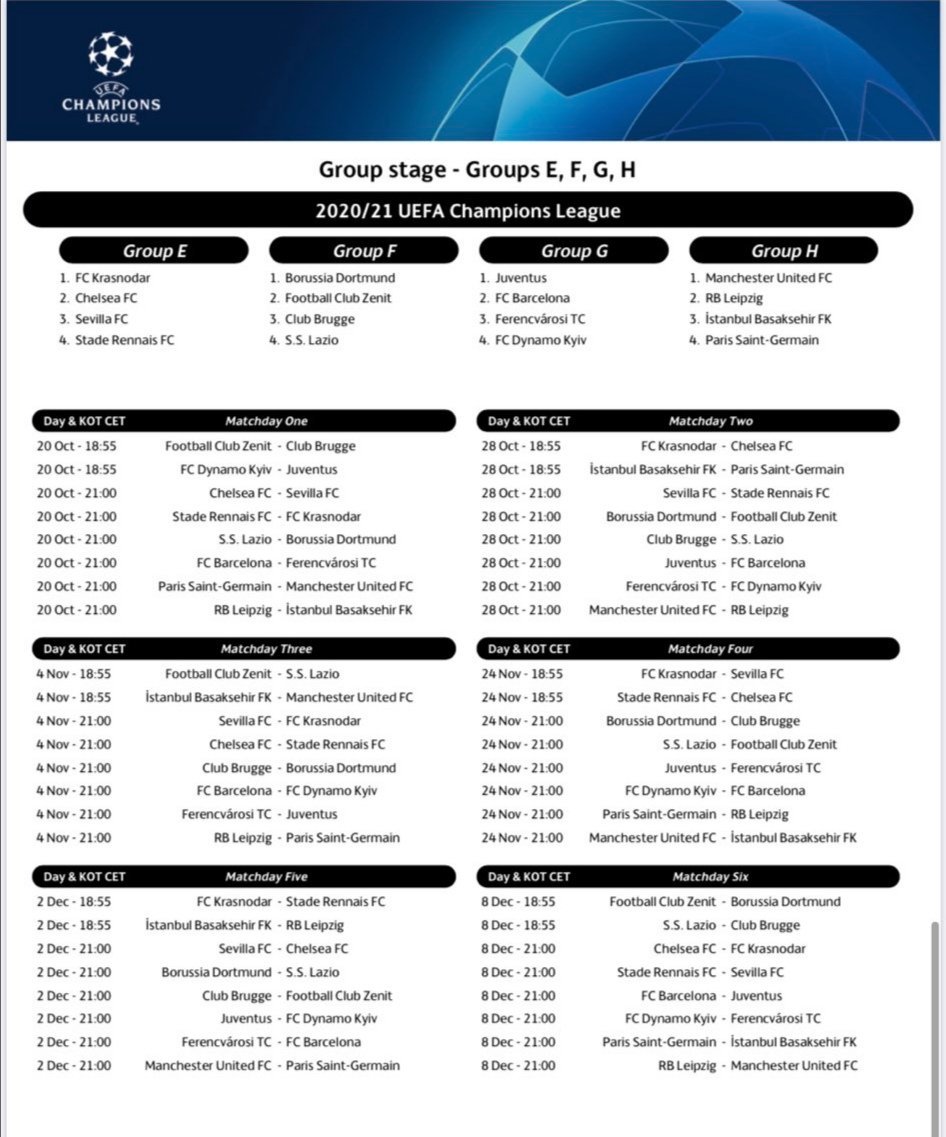
Champions League 2020/21 calendar: Real Madrid, Barcelona, Atletico Madrid and Sevilla fixtures | Marca

LIVESTREAM Ukrainian Championship: Kiev Capitals vs Lviv Lions, August 22, 16:30 CET (4:30 pm, 10:30 am ET)

Bandai Namco Esports on X: "Weekend is almost here which means we're opening our third week of #DBFZNC National Championship soon! As usual 🇪🇸 Spain will come first with some great matchups!














