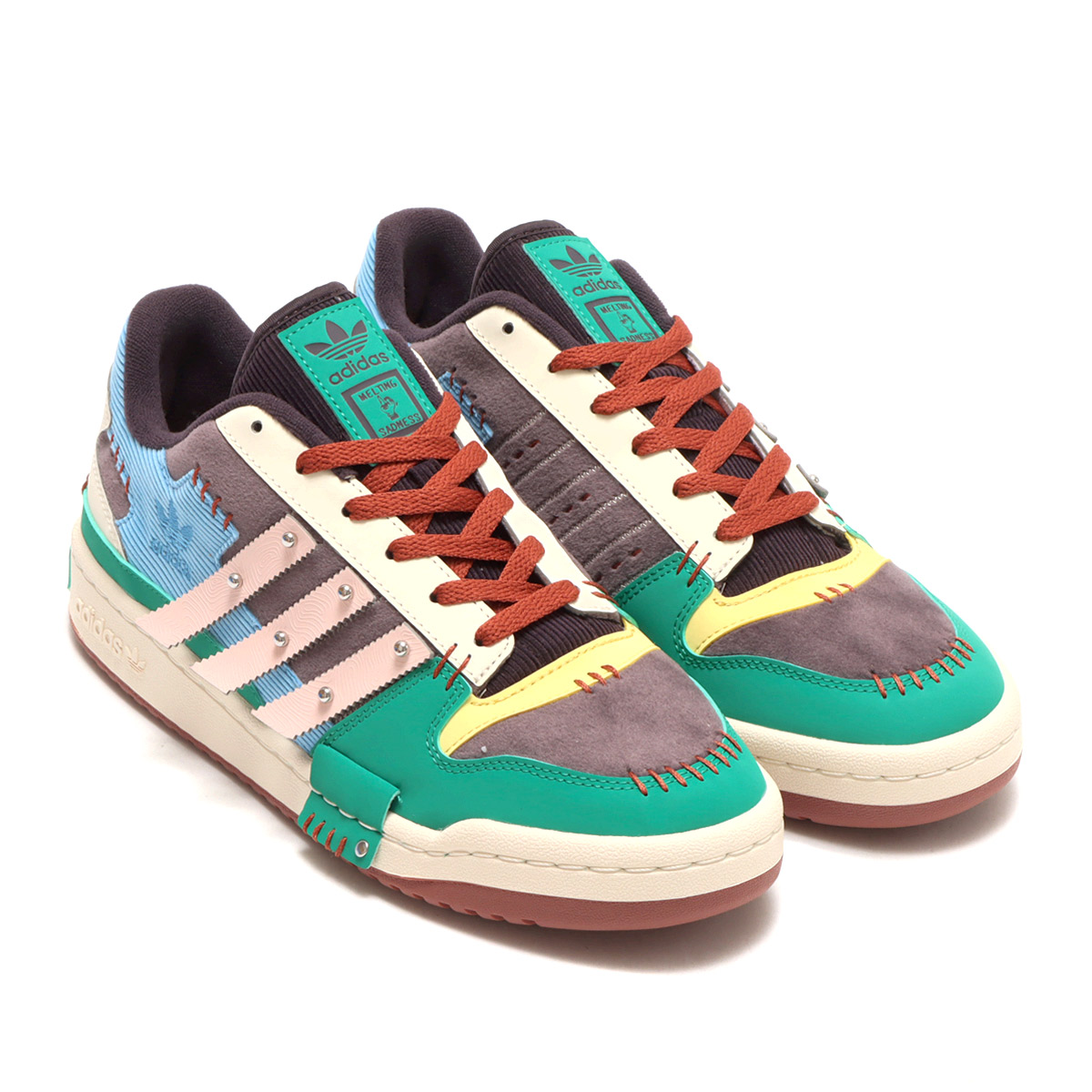
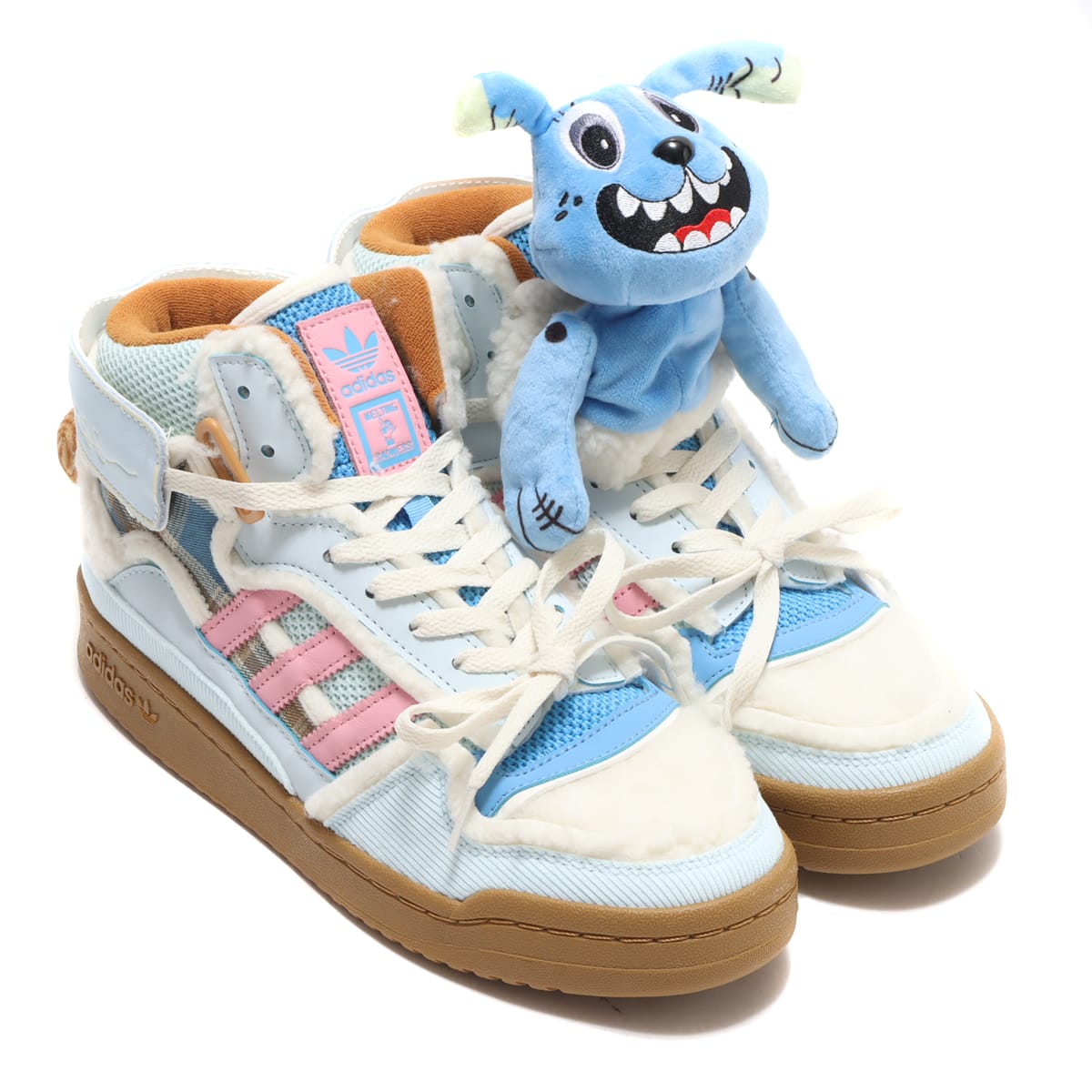
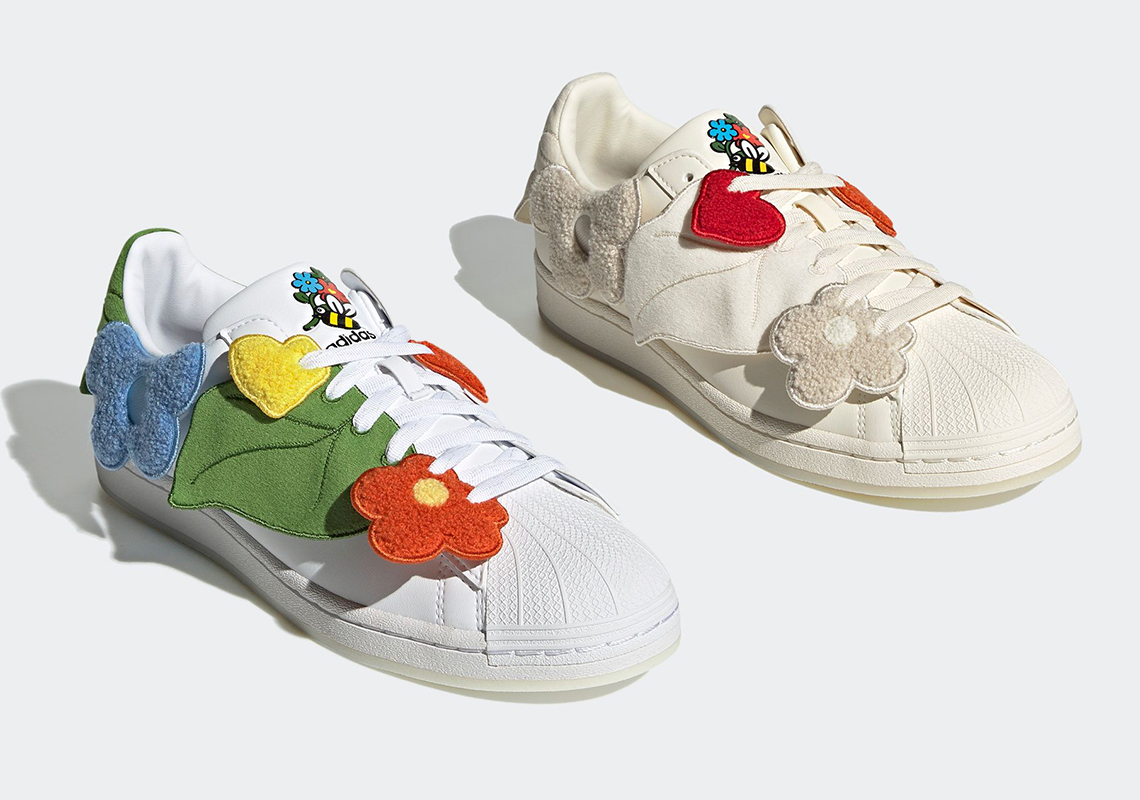
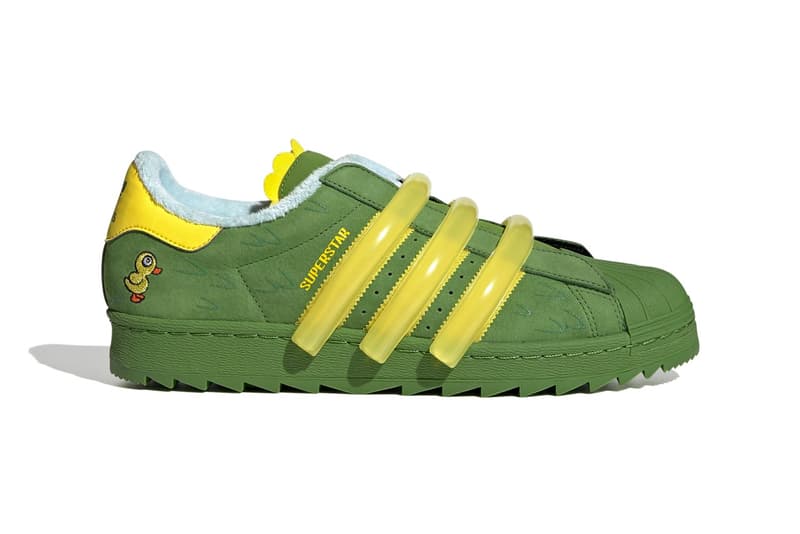
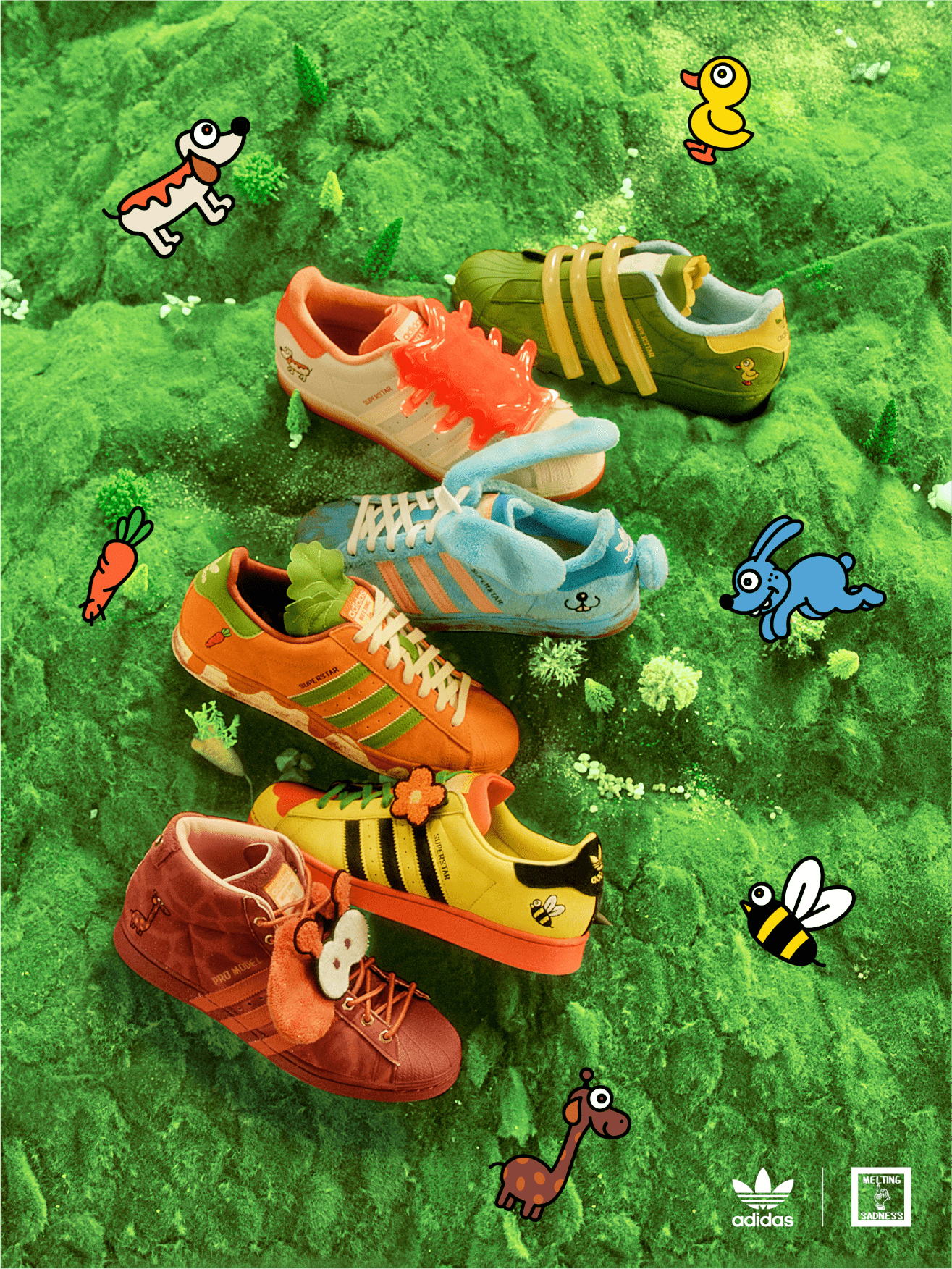
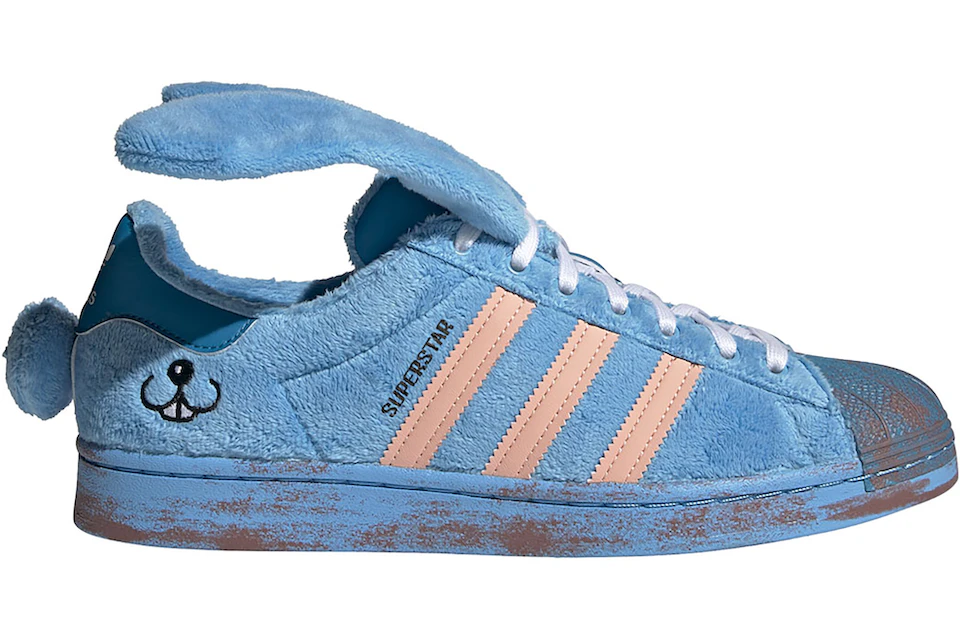
アディダス SST "クリア ブルー/エクルティント/オレンジ ラッシュ" adidas Superstar "Melting Sadness CNY Blue (2022)" - GY7012 - JP
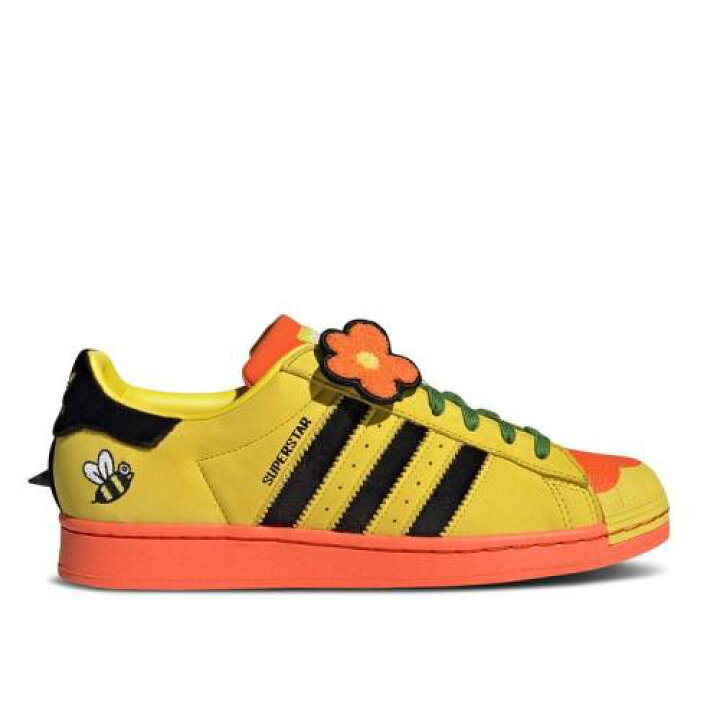

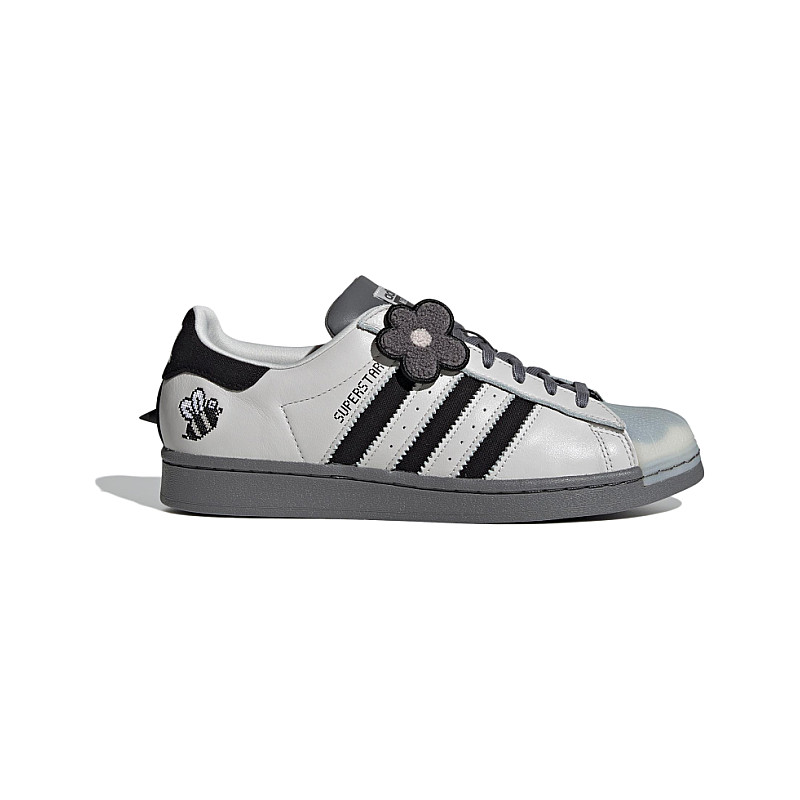
楽天市場】アディダス ADIDAS アディダス スーパースター 黄色 イエロー コア 黒色 ブラック 橙 オレンジ 'BEE YELLOW' スニーカー メンズ 【 SUPERSTAR YELLOW ORANGE ADIDAS MELTING SADNESS X WITH YOU PACK CORE BLACK SUPER : スニケス
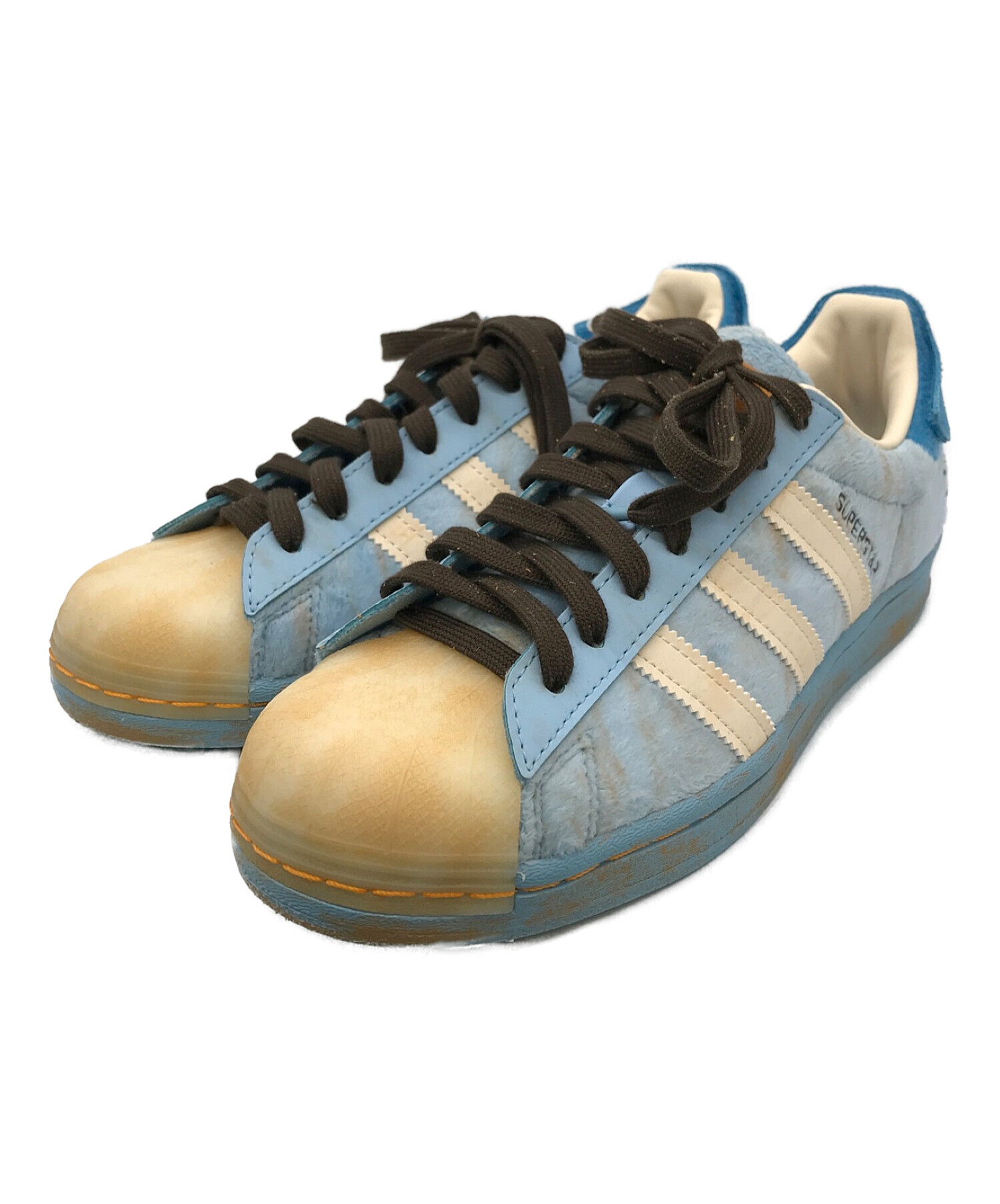

中古・古着通販】adidas×Melting Sadness SST (アディダス×メルティング サッドネス エスエスティー) SUPERSTAR ブルー サイズ:25|ブランド・古着通販 トレファク公式【TREFAC FASHION】

Adidas Forum X Melting Sadness in Accra Metropolitan - Shoes, Stone Unisex Collections | Jiji.com.gh

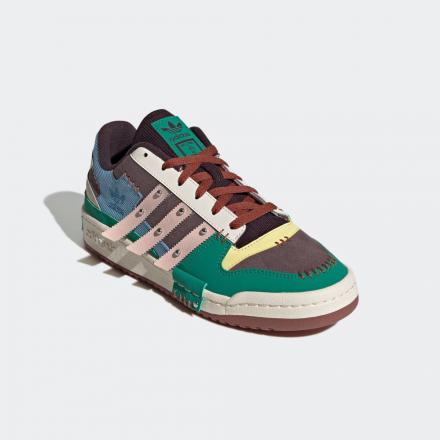
未使用品 アディダスオリジナルス adidas originals adidas originals×Melting Sadness アディダス オリジナルス×メルティング サッドネス GW8724 FORUM EXHIBIT LOW フォーラム エギジビット ロー スニーカー 28cm 019-202211210137 | ベクトルパーク
Melting Sadness × adidas originals Forum Low Core Black 25cm :sn-GW8726-25:SNEAKER SELECTION U-PICK - 通販 - Yahoo!ショッピング

MELTING SADNESS × ADIDAS ORIGINALS SUPERSTAR スーパースター (adidas/スニーカー) FZ5253 MELTING SADNESS ADIDAS ORIGINALS SUPERSTAR【BUYMA】

MELTING SADNESS x ADIDAS ORIGINAL FORUM LOW DOG/メルティング サッドネス x アディダス オリジナルス フォーラム ロー DOG GW8927 | スニーカーラボ